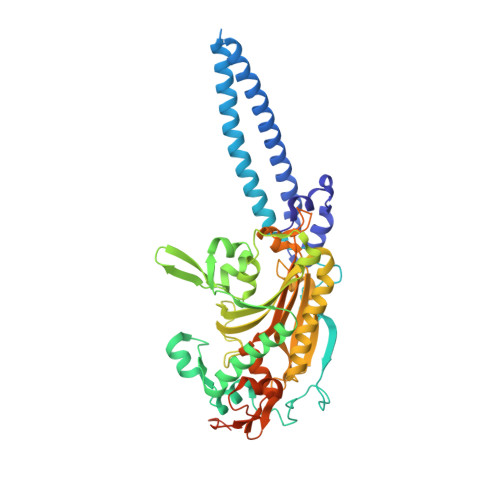
Unveiling the structural basis for translational ambiguity tolerance in a human fungal pathogen.
Rocha, R., Pereira, P.J., Santos, M.A., Macedo-Ribeiro, S.(2011) Proc Natl Acad Sci U S A 108: 14091-14096
- PubMed: 21825144
- DOI: https://doi.org/10.1073/pnas.1102835108
- Primary Citation of Related Structures:
3QNE, 3QO5, 3QO7, 3QO8 - PubMed Abstract:
In a restricted group of opportunistic fungal pathogens the universal leucine CUG codon is translated both as serine (97%) and leucine (3%), challenging the concept that translational ambiguity has a negative impact in living organisms. To elucidate the molecular mechanisms underlying the in vivo tolerance to a nonconserved genetic code alteration, we have undertaken an extensive structural analysis of proteins containing CUG-encoded residues and solved the crystal structures of the two natural isoforms of Candida albicans seryl-tRNA synthetase. We show that codon reassignment resulted in a nonrandom genome-wide CUG redistribution tailored to minimize protein misfolding events induced by the large-scale leucine-to-serine replacement within the CTG clade. Leucine or serine incorporation at the CUG position in C. albicans seryl-tRNA synthetase induces only local structural changes and, although both isoforms display tRNA serylation activity, the leucine-containing isoform is more active. Similarly, codon ambiguity is predicted to shape the function of C. albicans proteins containing CUG-encoded residues in functionally relevant positions, some of which have a key role in signaling cascades associated with morphological changes and pathogenesis. This study provides a first detailed analysis on natural reassignment of codon identity, unveiling a highly dynamic evolutionary pattern of thousands of fungal CUG codons to confer an optimized balance between protein structural robustness and functional plasticity.
Organizational Affiliation:
Instituto de Biologia Molecular e Celular, Universidade do Porto, 4150-180 Porto, Portugal.