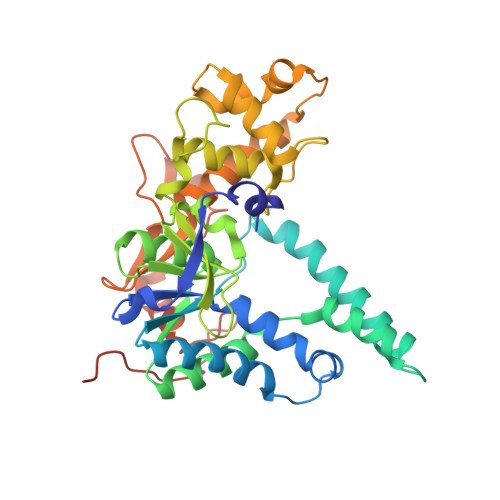
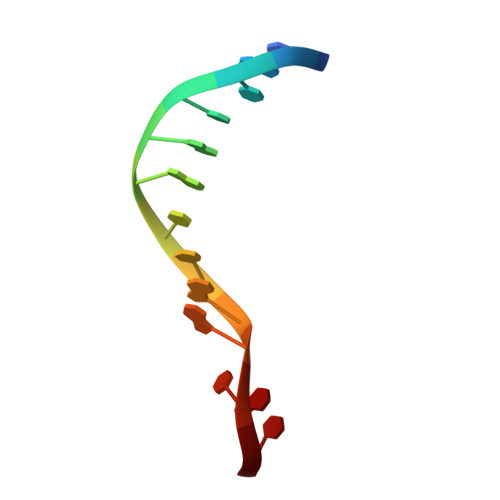
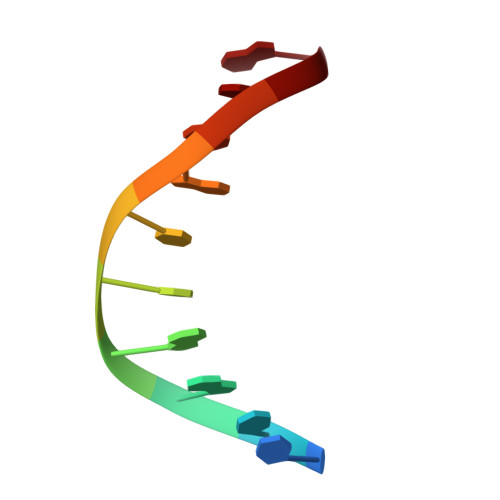
Structures of human exonuclease 1 DNA complexes suggest a unified mechanism for nuclease family.
Orans, J., McSweeney, E.A., Iyer, R.R., Hast, M.A., Hellinga, H.W., Modrich, P., Beese, L.S.(2011) Cell 145: 212-223
- PubMed: 21496642
- DOI: https://doi.org/10.1016/j.cell.2011.03.005
- Primary Citation of Related Structures:
3QE9, 3QEA, 3QEB - PubMed Abstract:
Human exonuclease 1 (hExo1) plays important roles in DNA repair and recombination processes that maintain genomic integrity. It is a member of the 5' structure-specific nuclease family of exonucleases and endonucleases that includes FEN-1, XPG, and GEN1. We present structures of hExo1 in complex with a DNA substrate, followed by mutagenesis studies, and propose a common mechanism by which this nuclease family recognizes and processes diverse DNA structures. hExo1 induces a sharp bend in the DNA at nicks or gaps. Frayed 5' ends of nicked duplexes resemble flap junctions, unifying the mechanisms of endo- and exonucleolytic processing. Conformational control of a mobile region in the catalytic site suggests a mechanism for allosteric regulation by binding to protein partners. The relative arrangement of substrate binding sites in these enzymes provides an elegant solution to a complex geometrical puzzle of substrate recognition and processing.
Organizational Affiliation:
Department of Biochemistry, Duke University Medical Center, Durham, NC 27710, USA.