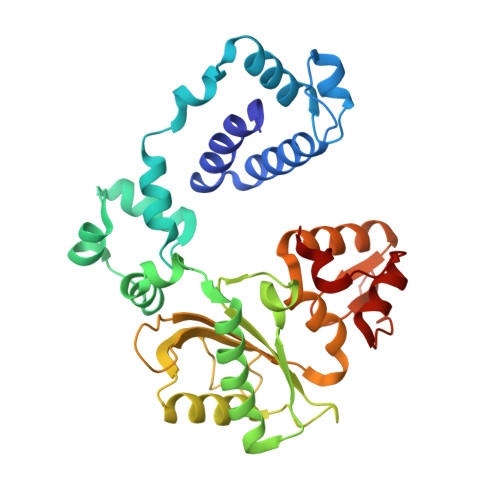
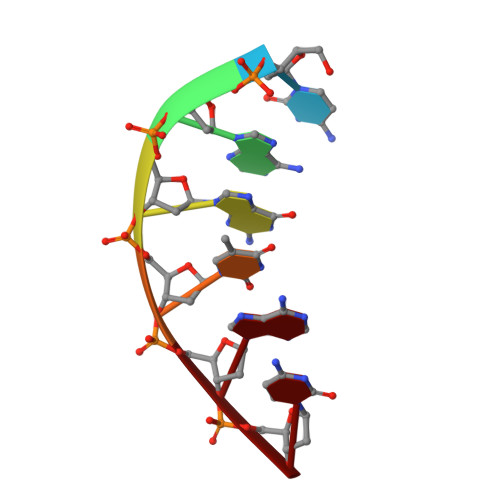
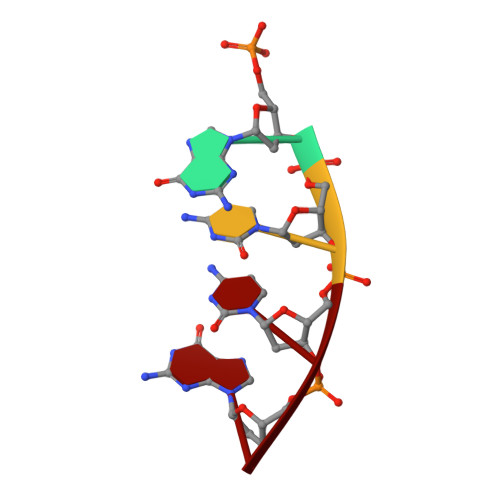
Interaction between DNA Polymerase lambda and anticancer nucleoside analogs.
Garcia-Diaz, M., Murray, M.S., Kunkel, T.A., Chou, K.M.(2010) J Biol Chem 285: 16874-16879
- PubMed: 20348107
- DOI: https://doi.org/10.1074/jbc.M109.094391
- Primary Citation of Related Structures:
3MDA, 3MDC - PubMed Abstract:
The anticancer activity of cytarabine (AraC) and gemcitabine (dFdC) is thought to result from chain termination after incorporation into DNA. To investigate their incorporation into DNA at atomic level resolution, we present crystal structures of human DNA polymerase lambda (Pol lambda) bound to gapped DNA and containing either AraC or dFdC paired opposite template dG. These structures reveal that AraC and dFdC can bind within the nascent base pair binding pocket of Pol lambda. Although the conformation of the ribose of AraCTP is similar to that of normal dCTP, the conformation of dFdCTP is significantly different. Consistent with these structures, Pol lambda efficiently incorporates AraCTP but not dFdCTP. The data are consistent with the possibility that Pol lambda could modulate the cytotoxic effect of AraC.
Organizational Affiliation:
Laboratory of Structural Biology and Laboratory of Molecular Genetics, Department of Health and Human Services, NIEHS, National Institutes of Health, Research Triangle Park, North Carolina 27709, USA.