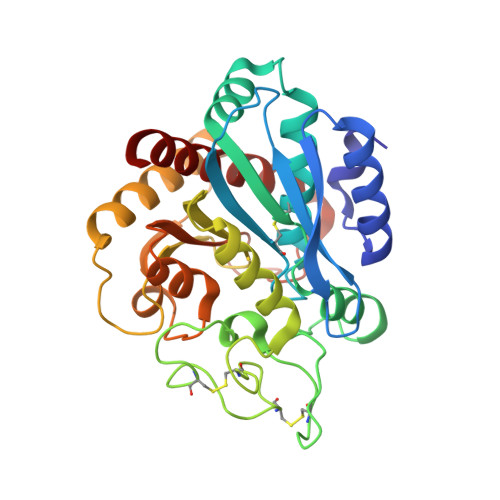
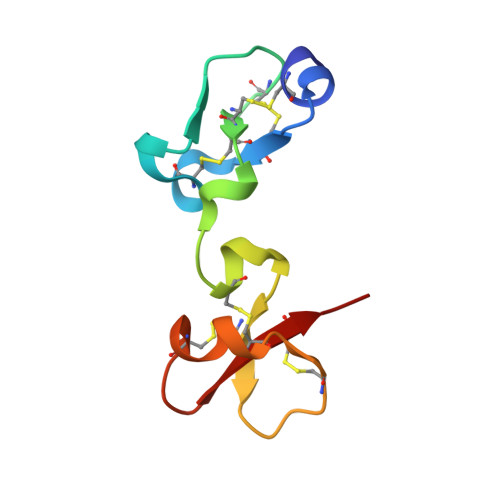
Insights into the molecular inactivation mechanism of human activated thrombin-activatable fibrinolysis inhibitor
Sanglas, L., Arolas, J.L., Valnickova, Z., Aviles, F.X., Enghild, J.J., Gomis-Ruth, F.X.(2010) J Thromb Haemost 8: 1056-1065
- PubMed: 20088943
- DOI: https://doi.org/10.1111/j.1538-7836.2010.03740.x
- Primary Citation of Related Structures:
3LMS - PubMed Abstract:
Thrombin-activatable fibrinolysis inhibitor (TAFI) is a validated target for thrombotic diseases. TAFI is converted in vivo to activated TAFI (TAFIa) by removal of its pro-domain. Whereas TAFI is stable and persists in the circulation, possibly in complex with plasminogen, TAFIa is unstable and poorly soluble, with a half-life of minutes.
Organizational Affiliation:
Departament de Bioquímica i Biologia Molecular, Facultat de Ciències, Institut de Biotecnologia i de Biomedicina, Universitat Autònoma de Barcelona, Bellaterra, Spain.