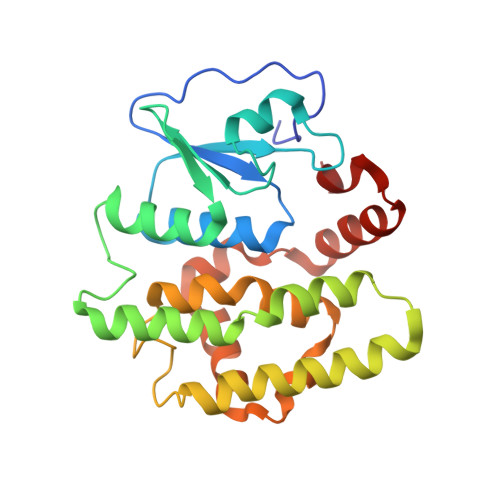
Novel folding and stability defects cause a deficiency of human glutathione transferase omega 1.
Zhou, H., Brock, J., Casarotto, M.G., Oakley, A.J., Board, P.G.(2011) J Biol Chem 286: 4271-4279
- PubMed: 21106529
- DOI: https://doi.org/10.1074/jbc.M110.197822
- Primary Citation of Related Structures:
3LFL - PubMed Abstract:
The polymorphic deletion of Glu-155 from human glutathione transferase omega1 (GSTO1-1) occurs in most populations. Although the recombinant ΔGlu-155 enzyme expressed in Escherichia coli is active, the deletion causes a deficiency of the active enzyme in vivo. The crystal structure and the folding/unfolding kinetics of the ΔGlu-155 variant were determined in order to investigate the cause of the rapid loss of the enzyme in human cells. The crystal structure revealed altered packing around the Glu-155 deletion, an increase in the predicted solvent-accessible area and a corresponding reduction in the buried surface area. This increase in solvent accessibility was consistent with an elevated Stern-Volmer constant. The unfolding of both the wild type and ΔGlu-155 enzyme in urea is best described by a three-state model, and there is evidence for the more pronounced population of an intermediate state by the ΔGlu-155 enzymes. Studies using intrinsic fluorescence revealed a free energy change around 14.4 kcal/mol for the wild type compared with around 8.6 kcal/mol for the ΔGlu-155 variant, which indicates a decrease in stability associated with the Glu-155 deletion. Urea induced unfolding of the wild type GSTO1-1 was reversible through an initial fast phase followed by a second slow phase. In contrast, the ΔGlu-155 variant lacks the slow phase, indicating a refolding defect. It is possible that in some conditions in vivo, the increased solvent-accessible area and the low stability of the ΔGlu-155 variant may promote its unfolding, whereas the refolding defect limits its refolding, resulting in GSTO1-1 deficiency.
Organizational Affiliation:
John Curtin School of Medical Research, Australian National University, Canberra, Australian Capital Territory 2601, Australia.