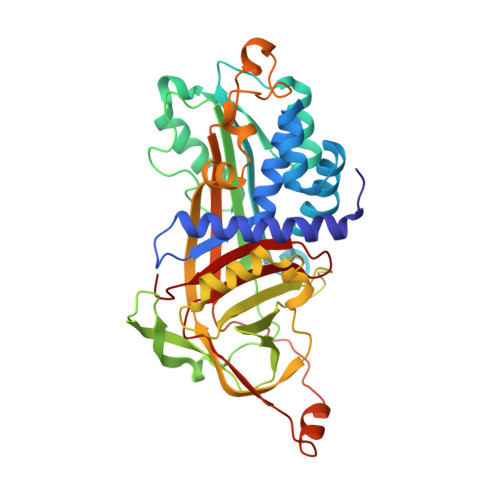
Arabidopsis AtSerpin1, crystal structure and in vivo interaction with its target protease RESPONSIVE TO DESICCATION-21 (RD21).
Lampl, N., Budai-Hadrian, O., Davydov, O., Joss, T.V., Harrop, S.J., Curmi, P.M., Roberts, T.H., Fluhr, R.(2010) J Biol Chem 285: 13550-13560
- PubMed: 20181955
- DOI: https://doi.org/10.1074/jbc.M109.095075
- Primary Citation of Related Structures:
3LE2 - PubMed Abstract:
In animals, protease inhibitors of the serpin family are associated with many physiological processes, including blood coagulation and innate immunity. Serpins feature a reactive center loop (RCL), which displays a protease target sequence as a bait. RCL cleavage results in an irreversible, covalent serpin-protease complex. AtSerpin1 is an Arabidopsis protease inhibitor that is expressed ubiquitously throughout the plant. The x-ray crystal structure of recombinant AtSerpin1 in its native stressed conformation was determined at 2.2 A. The electrostatic surface potential below the RCL was found to be highly positive, whereas the breach region critical for RCL insertion is an unusually open structure. AtSerpin1 accumulates in plants as a full-length and a cleaved form. Fractionation of seedling extracts by nonreducing SDS-PAGE revealed the presence of an additional slower migrating complex that was absent when leaves were treated with the specific cysteine protease inhibitor L-trans-epoxysuccinyl-L-leucylamido (4-guanidino)butane. Significantly, RESPONSIVE TO DESICCATION-21 (RD21) was the major protease labeled with the L-trans-epoxysuccinyl-L-leucylamido (4-guanidino)butane derivative DCG-04 in wild type extracts but not in extracts of mutant plants constitutively overexpressing AtSerpin1, indicating competition. Fractionation by nonreducing SDS-PAGE followed by immunoblotting with RD21-specific antibody revealed that the protease accumulated both as a free enzyme and in a complex with AtSerpin1. Importantly, both RD21 and AtSerpin1 knock-out mutants lacked the serpin-protease complex. The results establish that the major Arabidopsis plant serpin interacts with RD21. This is the first report of the structure and in vivo interaction of a plant serpin with its target protease.
Organizational Affiliation:
Department of Plant Sciences, Weizmann Institute of Science, Rehovot 76100, Israel.