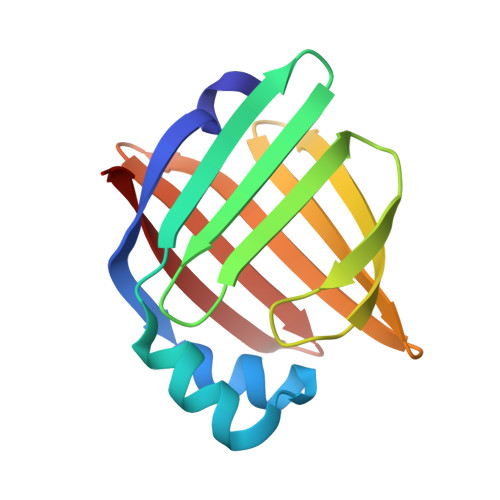
X-ray crystallographic analysis of adipocyte fatty acid binding protein (aP2) modified with 4-hydroxy-2-nonenal.
Hellberg, K., Grimsrud, P.A., Kruse, A.C., Banaszak, L.J., Ohlendorf, D.H., Bernlohr, D.A.(2010) Protein Sci 19: 1480-1489
- PubMed: 20509169
- DOI: https://doi.org/10.1002/pro.427
- Primary Citation of Related Structures:
3JS1, 3JSQ - PubMed Abstract:
Fatty acid binding proteins (FABP) have been characterized as facilitating the intracellular solubilization and transport of long-chain fatty acyl carboxylates via noncovalent interactions. More recent work has shown that the adipocyte FABP is also covalently modified in vivo on Cys117 with 4-hydroxy-2-nonenal (4-HNE), a bioactive aldehyde linked to oxidative stress and inflammation. To evaluate 4-HNE binding and modification, the crystal structures of adipocyte FABP covalently and noncovalently bound to 4-HNE have been solved to 1.9 A and 2.3 A resolution, respectively. While the 4-HNE in the noncovalently modified protein is coordinated similarly to a carboxylate of a fatty acid, the covalent form show a novel coordination through a water molecule at the polar end of the lipid. Other defining features between the two structures with 4-HNE and previously solved structures of the protein include a peptide flip between residues Ala36 and Lys37 and the rotation of the side chain of Phe57 into its closed conformation. Representing the first structure of an endogenous target protein covalently modified by 4-HNE, these results define a new class of in vivo ligands for FABPs and extend their physiological substrates to include bioactive aldehydes.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota-Twin Cities, Minneapolis, Minnesota, 55455, USA.