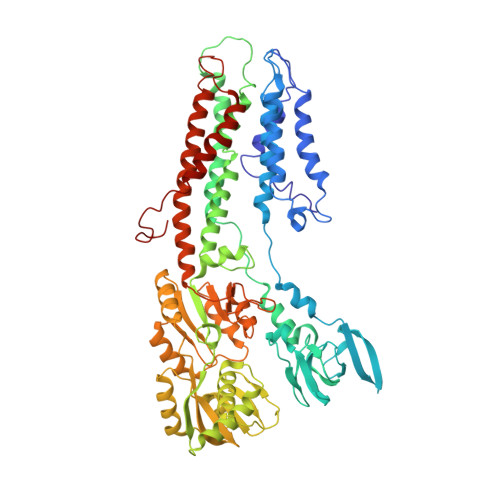
The Architecture of CopA from Archeaoglobus fulgidus Studied by Cryo-Electron Microscopy and Computational Docking.
Allen, G.S., Wu, C.C., Cardozo, T., Stokes, D.L.(2011) Structure 19: 1219-1232
- PubMed: 21820315
- DOI: https://doi.org/10.1016/j.str.2011.05.014
- Primary Citation of Related Structures:
3J08, 3J09 - PubMed Abstract:
CopA uses ATP to pump Cu(+) across cell membranes. X-ray crystallography has defined atomic structures of several related P-type ATPases. We have determined a structure of CopA at 10 Å resolution by cryo-electron microscopy of a new crystal form and used computational molecular docking to study the interactions between the N-terminal metal-binding domain (NMBD) and other elements of the molecule. We found that the shorter-chain lipids used to produce these crystals are associated with movements of the cytoplasmic domains, with a novel dimer interface and with disordering of the NMBD, thus offering evidence for the transience of its interaction with the other cytoplasmic domains. Docking identified a binding site that matched the location of the NMBD in our previous structure by cryo-electron microscopy, allowing a more detailed view of its binding configuration and further support for its role in autoinhibition.
Organizational Affiliation:
Skirball Institute, New York University School of Medicine, New York, NY 10016, USA.