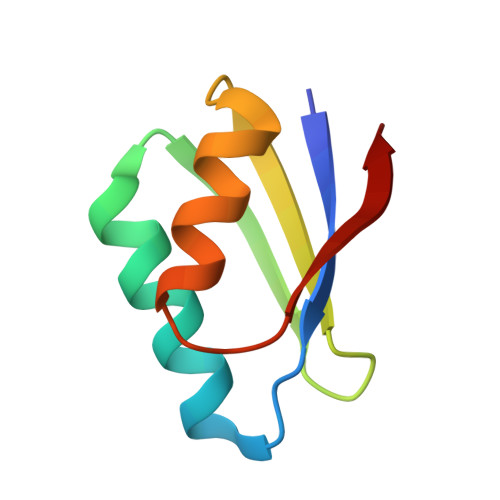
Crystal structures of cisplatin bound to a human copper chaperone.
Boal, A.K., Rosenzweig, A.C.(2009) J Am Chem Soc 131: 14196-14197
- PubMed: 19807176
- DOI: https://doi.org/10.1021/ja906363t
- Primary Citation of Related Structures:
3IWL, 3IWX - PubMed Abstract:
Copper trafficking proteins, including the chaperone Atox1 and the P(1B)-type ATPase ATP7B, have been implicated in cellular resistance to the anticancer drug cisplatin. We have determined two crystal structures of cisplatin-Atox1 adducts that reveal platinum coordination by the conserved CXXC copper-binding motif. Direct interaction of cisplatin with this functionally relevant site has significant implications for understanding the molecular basis for resistance mediated by copper transport pathways.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, Illinois 60208, USA.