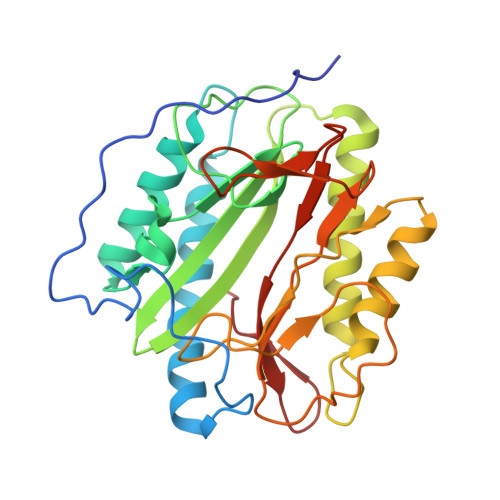
Catalysis and Inhibition of Mycobacterium tuberculosis Methionine Aminopeptidase
Lu, J.P., Chai, S.C., Ye, Q.Z.(2010) J Med Chem 53: 1329-1337
- PubMed: 20038112
- DOI: https://doi.org/10.1021/jm901624n
- Primary Citation of Related Structures:
3IU7, 3IU8, 3IU9 - PubMed Abstract:
Methionine aminopeptidase (MetAP) carries out an important cotranslational N-terminal methionine excision of nascent proteins and represents a potential target to develop antibacterial and antitubercular drugs. We cloned one of the two MetAPs in Mycobacterium tuberculosis (MtMetAP1c from the mapB gene) and purified it to homogeneity as an apoenzyme. Its activity required a divalent metal ion, and Co(II), Ni(II), Mn(II), and Fe(II) were among activators of the enzyme. Co(II) and Fe(II) had the tightest binding, while Ni(II) was the most efficient cofactor for the catalysis. MtMetAP1c was also functional in E. coli cells because a plasmid-expressed MtMetAP1c complemented the essential function of MetAP in E. coli and supported the cell growth. A set of potent MtMetAP1c inhibitors were identified, and they showed high selectivity toward the Fe(II)-form, the Mn(II)-form, or the Co(II) and Ni(II) forms of the enzyme, respectively. These metalloform selective inhibitors were used to assign the metalloform of the cellular MtMetAP1c. The fact that only the Fe(II)-form selective inhibitors inhibited the cellular MtMetAP1c activity and inhibited the MtMetAP1c-complemented cell growth suggests that Fe(II) is the native metal used by MtMetAP1c in an E. coli cellular environment. Finally, X-ray structures of MtMetAP1c in complex with three metalloform-selective inhibitors were analyzed, which showed different binding modes and different interactions with metal ions and active site residues.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, 635 Barnhill Drive, Indianapolis, Indiana 46202, USA.