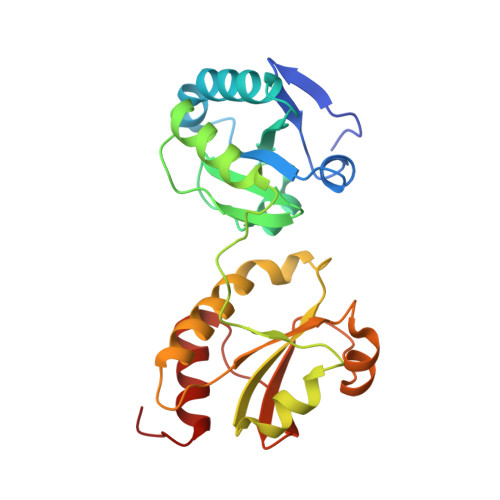
Structure of the Catalytic a(0)a Fragment of the Protein Disulfide Isomerase ERp72.
Kozlov, G., Azeroual, S., Rosenauer, A., Maattanen, P., Denisov, A.Y., Thomas, D.Y., Gehring, K.(2010) J Mol Biol 401: 618-625
- PubMed: 20600112
- DOI: https://doi.org/10.1016/j.jmb.2010.06.045
- Primary Citation of Related Structures:
3IDV - PubMed Abstract:
Protein disulfide isomerases (PDIs) are responsible for catalyzing the proper oxidation and isomerization of disulfide bonds of newly synthesized proteins in the endoplasmic reticulum (ER). The ER contains many different PDI-like proteins. Some, such as PDI, are general enzymes that directly recognize misfolded proteins while others, such as ERp57 and ERp72, have more specialized roles. Here, we report the high-resolution X-ray crystal structure of the N-terminal portion of ERp72 (also known as CaBP2 or PDI A4), which contains two a(0)a catalytic thioredoxin-like domains. The structure shows that the a(0) domain contains an additional N-terminal beta-strand and a different conformation of the beta5-alpha4 loop relative to other thioredoxin-like domains. The structure of the a domain reveals that a conserved arginine residue inserts into the hydrophobic core and makes a salt bridge with a conserved glutamate residue in the vicinity of the catalytic site. A structural model of full-length ERp72 shows that all three catalytic sites roughly face each other and positions the adjacent hydrophobic patches that are likely involved in protein substrate binding.
Organizational Affiliation:
Department of Biochemistry, Groupe de recherche axé sur la structure des protéines, McGill University, 3649 Promenade Sir William Osler, Montréal, Québec, Canada H3G 0B1.