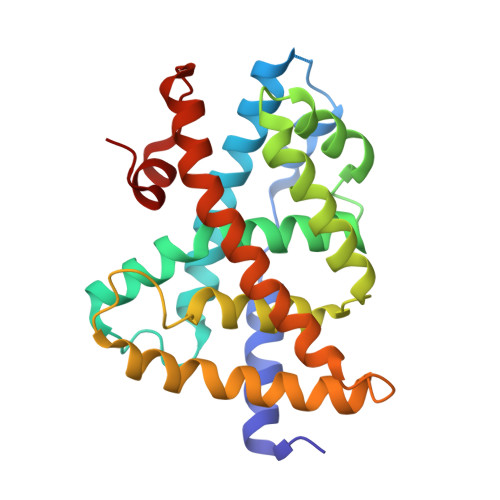
Substituted isoxazole analogs of farnesoid X receptor (FXR) agonist GW4064.
Bass, J.Y., Caldwell, R.D., Caravella, J.A., Chen, L., Creech, K.L., Deaton, D.N., Madauss, K.P., Marr, H.B., McFadyen, R.B., Miller, A.B., Parks, D.J., Todd, D., Williams, S.P., Wisely, G.B.(2009) Bioorg Med Chem Lett 19: 2969-2973
- PubMed: 19410460
- DOI: https://doi.org/10.1016/j.bmcl.2009.04.047
- Primary Citation of Related Structures:
3GD2 - PubMed Abstract:
Starting from the known FXR agonist GW 4064 1a, a series of alternately 3,5-substituted isoxazoles was prepared. Several of these analogs were potent full FXR agonists. A subset of this series, with a tether between the isoxazole ring and the 3-position aryl substituent, were equipotent FXR agonists to GW 4064 1a, with the 2,6-dimethyl phenol analog 1t having greater FRET FXR potency than GW 4064 1a.
Organizational Affiliation:
Department of Medicinal Chemistry, GlaxoSmithKline, Research Triangle Park, NC 27709, USA.