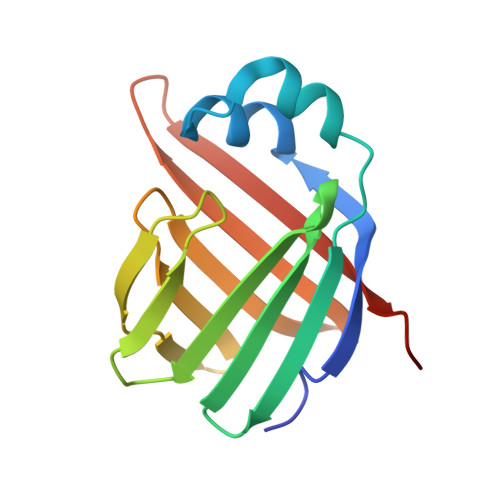
The X-Ray structure of zebrafish (Danio rerio) ileal bile acid-binding protein reveals the presence of binding sites on the surface of the protein molecule.
Capaldi, S., Saccomani, G., Fessas, D., Signorelli, M., Perduca, M., Monaco, H.L.(2009) J Mol Biol 385: 99-116
- PubMed: 18952094
- DOI: https://doi.org/10.1016/j.jmb.2008.10.007
- Primary Citation of Related Structures:
3ELX, 3ELZ, 3EM0 - PubMed Abstract:
The ileal bile acid-binding proteins (I-BABPs), also called ileal lipid-binding proteins or gastrotropins, belong to the family of the fatty acid-binding proteins and play an important role in the solubilization and transport of bile acids in the enterocyte. This article describes the expression, purification, crystallization, and three-dimensional structure determination of zebrafish (Danio rerio) I-BABP both in its apo form and bound to cholic acid. This is the first X-ray structure of an I-BABP. The structure of the apoprotein was determined to a resolution of 1.6 A, and two different monoclinic crystal forms of the holoprotein were solved and refined to 2.2 A resolution. Three protein molecules are present in the asymmetric unit of one of the co-crystal forms and two in the other, and therefore, the results of this study refer to observations made on five different protein molecules in the crystalline state. In every case, two cholate ligands were found bound in approximately the same position in the internal cavity of the protein molecules, but an unexpected result is the presence of clear and unambiguous electron density for several cholate molecules bound on hydrophobic patches on the surface of all the five independent protein molecules examined. Isothermal titration calorimetry was used for the thermodynamic characterization of the binding mechanism and has yielded results that are consistent with the X-ray data. Ligand binding is described in detail, and the conformational changes undergone by the protein molecule in the apo-to-holo transition are examined by superposition of the apo- and holoprotein models. The structure of the holoprotein is also compared with that of the liver BABP from the same species and those of other I-BABPs determined by NMR.
Organizational Affiliation:
Biocrystallography Laboratory, Department of Biotechnology, University of Verona, Verona, Italy.