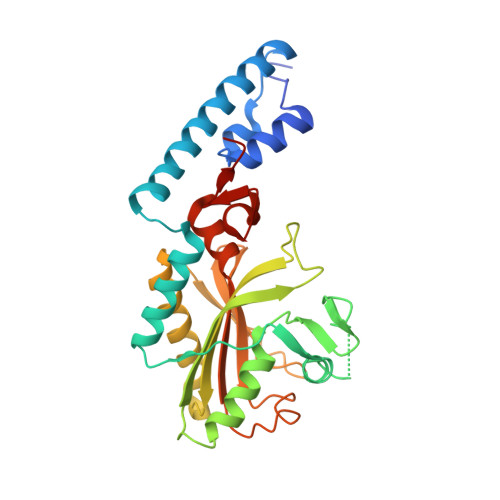
Structure of Desulfitobacterium hafniense PylSc, a pyrrolysyl-tRNA synthetase.
Lee, M.M., Jiang, R., Jain, R., Larue, R.C., Krzycki, J., Chan, M.K.(2008) Biochem Biophys Res Commun 374: 470-474
- PubMed: 18656445
- DOI: https://doi.org/10.1016/j.bbrc.2008.07.074
- Primary Citation of Related Structures:
3DSQ - PubMed Abstract:
Pyrrolysine, the 22nd genetically-encoded amino acid, is charged onto its specific tRNA by PylS, a pyrrolysyl-tRNA synthetase. While PylS is found as a single protein in certain archaeal methanogens, in the gram-positive bacterium Desulfitobacterium hafniense, PylS is divided into two separate proteins, PylSn and PylSc, corresponding to the N-terminal and C-terminal domains of the single PylS protein found in methanogens. Previous crystallographic studies have provided the structure of a truncated C-terminal portion of the archaeal Methanosarcina mazei PylS associated with catalysis. Here, we report the apo 2.1A resolution structure of the intact D. hafniense PylSc protein and compare it to structures of the C-terminal truncated PylS from methanogenic species. In PylSc, the hydrophobic pocket binding the ring of pyrrolysine is more constrained than in the archaeal enzyme; other structural differences are also apparent.
Organizational Affiliation:
The Ohio State Biophysics Program, The Ohio State University, 484 West 12th Avenue, Columbus, OH 43210, USA.