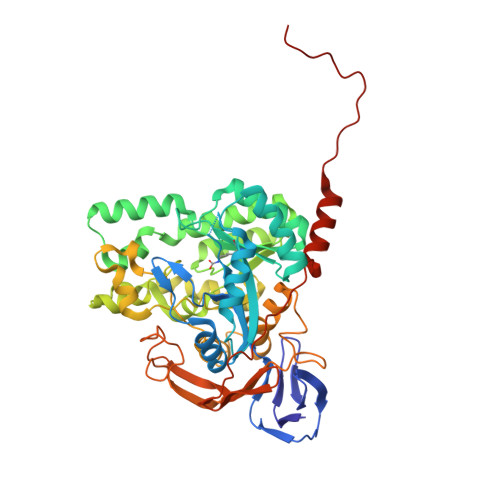
Structure of dihydropyrimidinase from Sinorhizobium meliloti CECT4114: new features in an amidohydrolase family member
Martinez-Rodriguez, S., Martinez-Gomez, A.I., Clemente-Jimenez, J.M., Rodriguez-Vico, F., Garcia-Ruiz, J.M., Las Heras-Vazquez, F.J., Gavira, J.A.(2010) J Struct Biol 169: 200-208
- PubMed: 19895890
- DOI: https://doi.org/10.1016/j.jsb.2009.10.013
- Primary Citation of Related Structures:
3DC8 - PubMed Abstract:
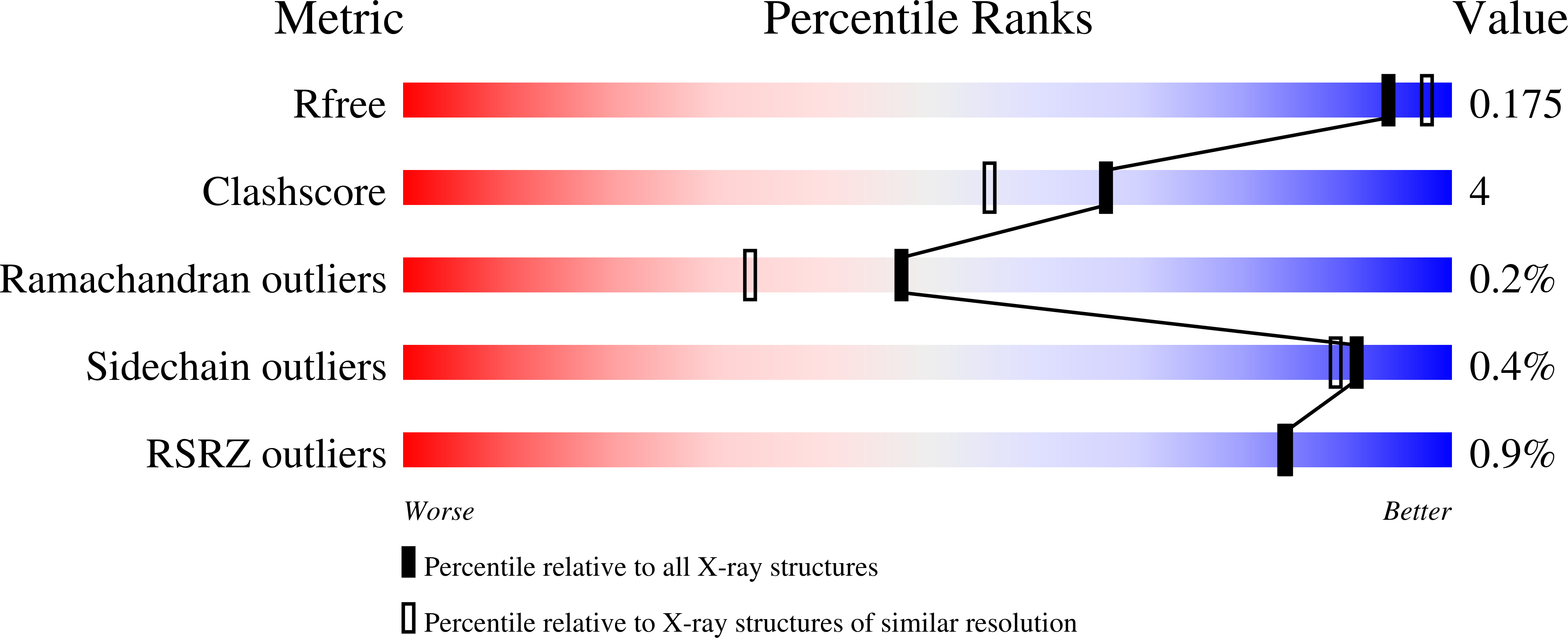
The recombinant dihydropyrimidinase from Sinorhizobium meliloti CECT4114 (SmelDhp) has been characterised and its crystal structure elucidated at 1.85A. The global architecture of the protein is reminiscent of that of the amidohydrolase superfamily, consisting of two domains; an (alpha/beta)(8) TIM-like barrel domain, where the catalytic centre is located, and a smaller beta-sheet sandwich domain of unknown function. The c-terminal tails of each subunit extend toward another monomer in a swapping-like manner, creating a hydrogen bond network which suggests its implication in protein oligomerisation. Mutational and structural evidence suggest the involvement of a conserved tyrosine in the reaction mechanism of the enzyme. SmelDhp presents both hydantoinase and dihydropyrimidinase activities, with higher affinity for the natural six-membered ring substrates. For the five-membered ring substrates, affinity was greater for those with aliphatic and apolar groups in the 5th carbon atom, with the highest rates of hydrolysis for d-5-methyl and d-5-ethyl hydantoin (k(cat)/K(m)=2736+/-380 and 944+/-52M(-1)s(-1), respectively). The optimal conditions for the enzyme activity were found to be 60 degrees C of temperature at pH 8.0. SmelDhp retains 95% of its activity after 6-hour preincubation at 60 degrees C. This is the first dihydropyrimidinase used for the hydrolytic opening of non-natural 6-monosubstituted dihydrouracils, which may be exploited for the production of beta-amino acids.
Organizational Affiliation:
Dpto. Química Física, Bioquímica y Química Inorgánica, Universidad de Almería, Almería, Spain. srodrig@ual.es