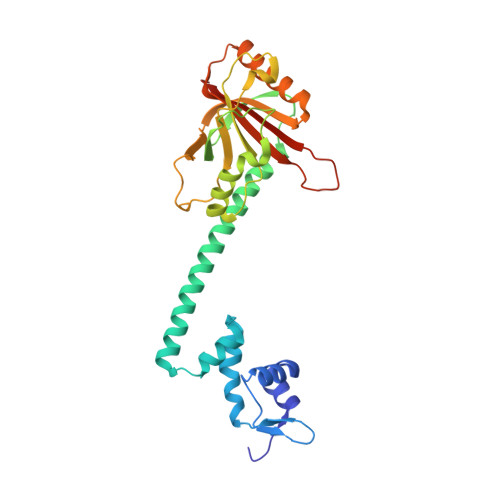
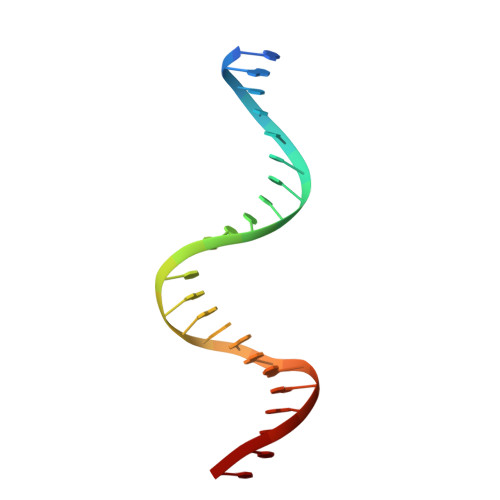
Structures of BmrR-Drug Complexes Reveal a Rigid Multidrug Binding Pocket and Transcription Activation through Tyrosine Expulsion
Newberry, K.J., Huffman, J.L., Miller, M.C., Vazquez-Laslop, N., Neyfakh, A.A., Brennan, R.G.(2008) J Biol Chem 283: 26795-26804
- PubMed: 18658145
- DOI: https://doi.org/10.1074/jbc.M804191200
- Primary Citation of Related Structures:
3D6Y, 3D6Z, 3D70, 3D71 - PubMed Abstract:
BmrR is a member of the MerR family and a multidrug binding transcription factor that up-regulates the expression of the bmr multidrug efflux transporter gene in response to myriad lipophilic cationic compounds. The structural mechanism by which BmrR binds these chemically and structurally different drugs and subsequently activates transcription is poorly understood. Here, we describe the crystal structures of BmrR bound to rhodamine 6G (R6G) or berberine (Ber) and cognate DNA. These structures reveal each drug stacks against multiple aromatic residues with their positive charges most proximal to the carboxylate group of Glu-253 and that, unlike other multidrug binding pockets, that of BmrR is rigid. Substitution of Glu-253 with either alanine (E253A) or glutamine (E253Q) results in unpredictable binding affinities for R6G, Ber, and tetraphenylphosphonium. Moreover, these drug binding studies reveal that the negative charge of Glu-253 is not important for high affinity binding to Ber and tetraphenylphosphonium but plays a more significant, but unpredictable, role in R6G binding. In vitro transcription data show that E253A and E253Q are constitutively active, and structures of the drug-free E253A-DNA and E253Q-DNA complexes support a transcription activation mechanism requiring the expulsion of Tyr-152 from the multidrug binding pocket. In sum, these data delineate the mechanism by which BmrR binds lipophilic, monovalent cationic compounds and suggest the importance of the redundant negative electrostatic nature of this rigid drug binding pocket that can be used to discriminate against molecules that are not substrates of the Bmr multidrug efflux pump.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of Texas M.D. Anderson Cancer Center, Houston, Texas 77030-4009, USA.