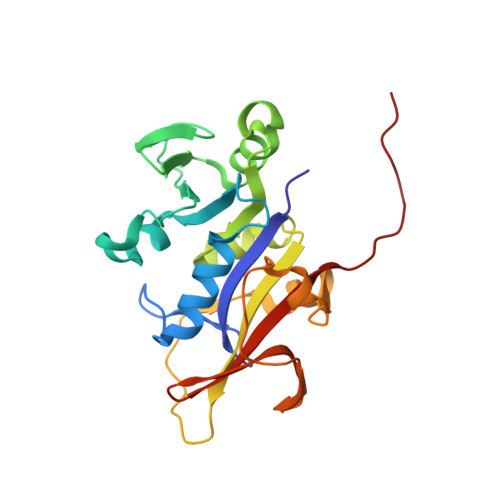
Structure-guided development of efficacious antifungal agents targeting Candida glabrata dihydrofolate reductase.
Liu, J., Bolstad, D.B., Smith, A.E., Priestley, N.D., Wright, D.L., Anderson, A.C.(2008) Chem Biol 15: 990-996
- PubMed: 18804036
- DOI: https://doi.org/10.1016/j.chembiol.2008.07.013
- Primary Citation of Related Structures:
3CSE - PubMed Abstract:
Candida glabrata is a lethal fungal pathogen resistant to many antifungal agents and has emerged as a critical target for drug discovery. Over the past several years, we have been developing a class of propargyl-linked antifolates as antimicrobials and hypothesized that these compounds could be effective inhibitors of dihydrofolate reductase (DHFR) from C. glabrata. We initially screened a small collection of these inhibitors and found modest levels of potency. Subsequently, we determined the crystal structure of C. glabrata DHFR bound to a representative inhibitor with data to 1.6 A resolution. Using this structure, we designed and synthesized second-generation inhibitors. These inhibitors bind the C. glabrata DHFR enzyme with subnanomolar potency, display greater than 2000-fold levels of selectivity over the human enzyme, and inhibit the growth of C. glabrata at levels observed with clinically employed therapeutics.
Organizational Affiliation:
Department of Pharmaceutical Sciences, University of Connecticut, 69 N. Eagleville Road, Storrs, CT 0626, USA.