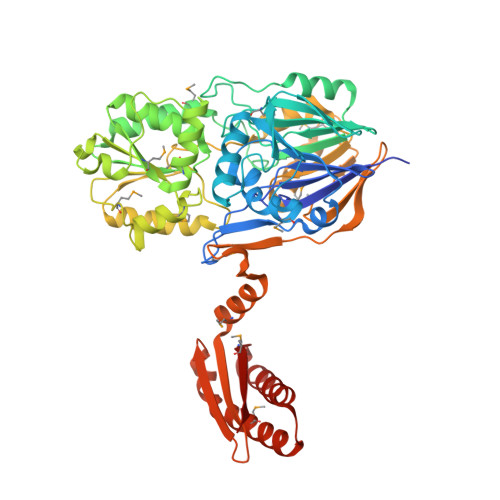
Structural insights into the dual activity of RNase J
de la Sierra-Gallay, I.L., Zig, L., Jamalli, A., Putzer, H.(2008) Nat Struct Mol Biol 15: 206-212
- PubMed: 18204464
- DOI: https://doi.org/10.1038/nsmb.1376
- Primary Citation of Related Structures:
3BK1, 3BK2 - PubMed Abstract:
The maturation and stability of RNA transcripts is controlled by a combination of endo- and exoRNases. RNase J is unique, as it combines an RNase E-like endoribonucleolytic and a 5'-to-3' exoribonucleolytic activity in a single polypeptide. The structural basis for this dual activity is unknown. Here we report the crystal structures of Thermus thermophilus RNase J and its complex with uridine 5'-monophosphate. A binding pocket coordinating the phosphate and base moieties of the nucleotide in the vicinity of the catalytic center provide a rationale for the 5'-monophosphate-dependent 5'-to-3' exoribonucleolytic activity. We show that this dependence is strict; an initial 5'-PPP transcript cannot be degraded exonucleolytically from the 5'-end. Our results suggest that RNase J might switch promptly from endo- to exonucleolytic mode on the same RNA, a property that has important implications for RNA metabolism in numerous prokaryotic organisms and plant organelles containing RNase J orthologs.
Organizational Affiliation:
FRC550, Institut de Biologie Physico-Chimique, 13 rue Pierre et Marie Curie, 75005 Paris, France.