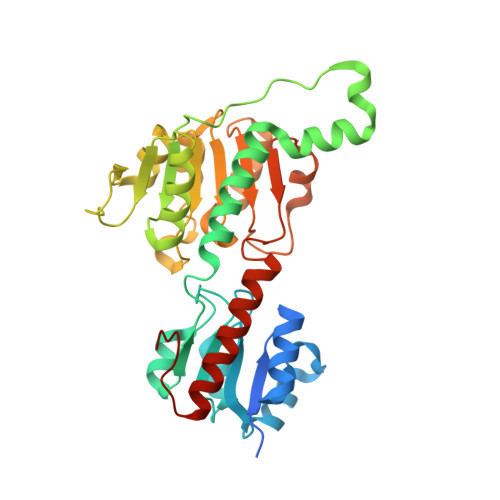
Structure and substrate docking of a hydroxy(phenyl)pyruvate reductase from the higher plant Coleus blumei Benth.
Janiak, V., Petersen, M., Zentgraf, M., Klebe, G., Heine, A.(2010) Acta Crystallogr D Biol Crystallogr 66: 593-603
- PubMed: 20445235
- DOI: https://doi.org/10.1107/S0907444910006360
- Primary Citation of Related Structures:
3BA1, 3BAZ - PubMed Abstract:
Hydroxy(phenyl)pyruvate reductase [H(P)PR] belongs to the family of D-isomer-specific 2-hydroxyacid dehydrogenases and catalyzes the reduction of hydroxyphenylpyruvates as well as hydroxypyruvate and pyruvate to the corresponding lactates. Other non-aromatic substrates are also accepted. NADPH is the preferred cosubstrate. The crystal structure of the enzyme from Coleus blumei (Lamiaceae) has been determined at 1.47 A resolution. In addition to the apoenzyme, the structure of a complex with NADP(+) was determined at a resolution of 2.2 A. H(P)PR is a dimer with a molecular mass of 34 113 Da per subunit. The structure is similar to those of other members of the enzyme family and consists of two domains separated by a deep catalytic cleft. To gain insights into substrate binding, several compounds were docked into the cosubstrate complex structure using the program AutoDock. The results show two possible binding modes with similar docking energy. However, only binding mode A provides the necessary environment in the active centre for hydride and proton transfer during reduction, leading to the formation of the (R)-enantiomer of lactate and/or hydroxyphenyllactate.
Organizational Affiliation:
Institut für Pharmazeutische Biologie, Philipps-Universität Marburg, Deutschhausstrasse 17A, 35037 Marburg, Germany.