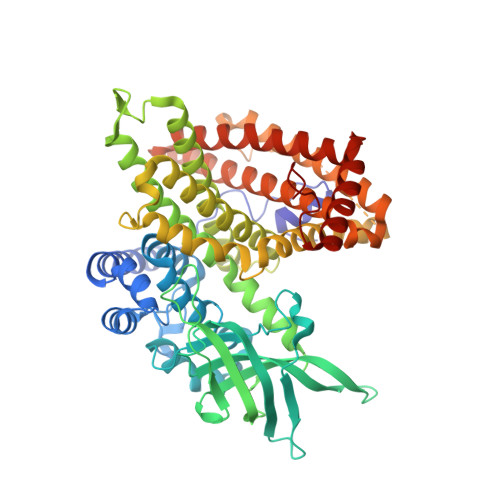
Structural basis for substrate fatty acyl chain specificity: crystal structure of human very-long-chain acyl-CoA dehydrogenase.
McAndrew, R.P., Wang, Y., Mohsen, A.W., He, M., Vockley, J., Kim, J.J.(2008) J Biol Chem 283: 9435-9443
- PubMed: 18227065
- DOI: https://doi.org/10.1074/jbc.M709135200
- Primary Citation of Related Structures:
3B96 - PubMed Abstract:
Very-long-chain acyl-CoA dehydrogenase (VLCAD) is a member of the family of acyl-CoA dehydrogenases (ACADs). Unlike the other ACADs, which are soluble homotetramers, VLCAD is a homodimer associated with the mitochondrial membrane. VLCAD also possesses an additional 180 residues in the C terminus that are not present in the other ACADs. We have determined the crystal structure of VLCAD complexed with myristoyl-CoA, obtained by co-crystallization, to 1.91-A resolution. The overall fold of the N-terminal approximately 400 residues of VLCAD is similar to that of the soluble ACADs including medium-chain acyl-CoA dehydrogenase (MCAD). The novel C-terminal domain forms an alpha-helical bundle that is positioned perpendicular to the two N-terminal helical domains. The fatty acyl moiety of the bound substrate/product is deeply imbedded inside the protein; however, the adenosine pyrophosphate portion of the C14-CoA ligand is disordered because of partial hydrolysis of the thioester bond and high mobility of the CoA moiety. The location of Glu-422 with respect to the C2-C3 of the bound ligand and FAD confirms Glu-422 to be the catalytic base. In MCAD, Gln-95 and Glu-99 form the base of the substrate binding cavity. In VLCAD, these residues are glycines (Gly-175 and Gly-178), allowing the binding channel to extend for an additional 12A and permitting substrate acyl chain lengths as long as 24 carbons to bind. VLCAD deficiency is among the more common defects of mitochondrial beta-oxidation and, if left undiagnosed, can be fatal. This structure allows us to gain insight into how a variant VLCAD genotype results in a clinical phenotype.
Organizational Affiliation:
Department of Biochemistry, Medical College of Wisconsin, 8701 Watertown Plank Road, Milwaukee, WI 53226, USA.