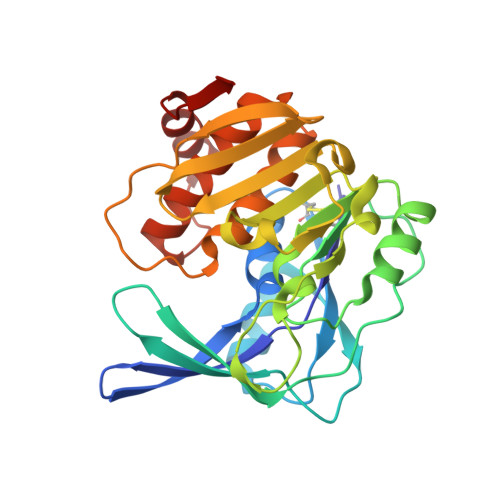
An uncharacterized member of the ribokinase family in Thermococcus kodakarensis exhibits myo-inositol kinase activity.
Sato, T., Fujihashi, M., Miyamoto, Y., Kuwata, K., Kusaka, E., Fujita, H., Miki, K., Atomi, H.(2013) J Biol Chem 288: 20856-20867
- PubMed: 23737529
- DOI: https://doi.org/10.1074/jbc.M113.457259
- Primary Citation of Related Structures:
3W4S - PubMed Abstract:
Here we performed structural and biochemical analyses on the TK2285 gene product, an uncharacterized protein annotated as a member of the ribokinase family, from the hyperthermophilic archaeon Thermococcus kodakarensis. The three-dimensional structure of the TK2285 protein resembled those of previously characterized members of the ribokinase family including ribokinase, adenosine kinase, and phosphofructokinase. Conserved residues characteristic of this protein family were located in a cleft of the TK2285 protein as in other members whose structures have been determined. We thus examined the kinase activity of the TK2285 protein toward various sugars recognized by well characterized ribokinase family members. Although activity with sugar phosphates and nucleosides was not detected, kinase activity was observed toward d-allose, d-lyxose, d-tagatose, d-talose, d-xylose, and d-xylulose. Kinetic analyses with the six sugar substrates revealed high Km values, suggesting that they were not the true physiological substrates. By examining activity toward amino sugars, sugar alcohols, and disaccharides, we found that the TK2285 protein exhibited prominent kinase activity toward myo-inositol. Kinetic analyses with myo-inositol revealed a greater kcat and much lower Km value than those obtained with the monosaccharides, resulting in over a 2,000-fold increase in kcat/Km values. TK2285 homologs are distributed among members of Thermococcales, and in most species, the gene is positioned close to a myo-inositol monophosphate synthase gene. Our results suggest the presence of a novel subfamily of the ribokinase family whose members are present in Archaea and recognize myo-inositol as a substrate.
Organizational Affiliation:
From the Department of Synthetic Chemistry and Biological Chemistry, Graduate School of Engineering, Kyoto University, Katsura, Nishikyo-ku, Kyoto 615-8510, Japan,; Core Research for Evolutional Science and Technology (CREST), Japan Science and Technology Agency (JST), 7 Gobancho, Chiyoda-ku, Tokyo 102-0076, Japan.