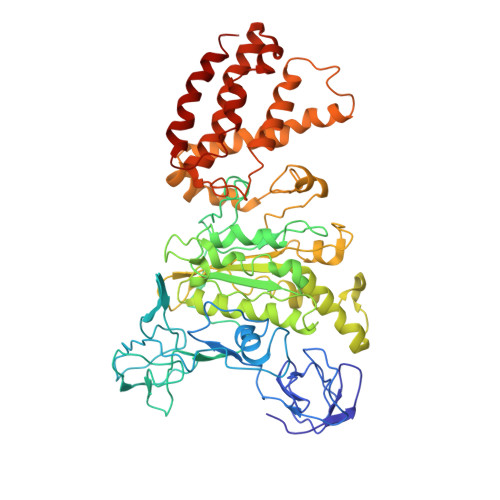
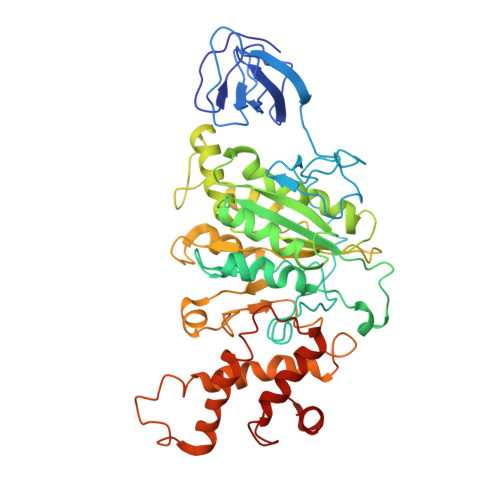
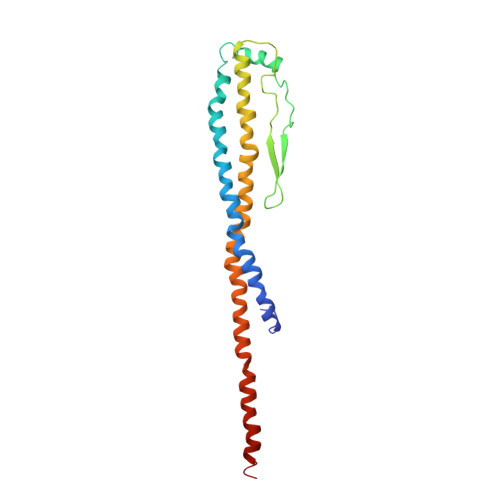
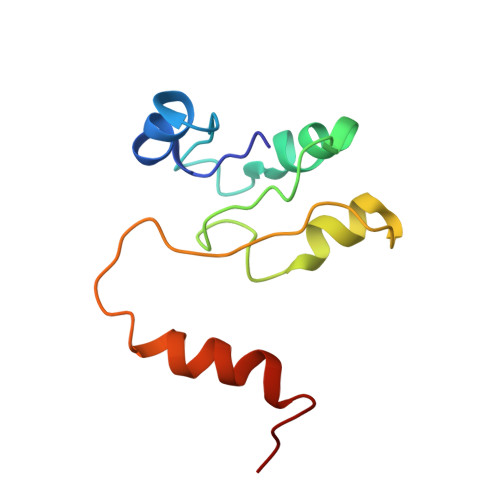
Origin of Asymmetry at the Intersubunit Interfaces of V1-ATPase from Thermusthermophilus
Nagamatsu, Y., Takeda, K., Kuranaga, T., Numoto, N., Miki, K.(2013) J Mol Biol 425: 2699-2708
- PubMed: 23639357
- DOI: https://doi.org/10.1016/j.jmb.2013.04.022
- Primary Citation of Related Structures:
3W3A - PubMed Abstract:
V-type ATPase (V-ATPase) is one of the rotary ATPase complexes that mediate energy conversion between the chemical energy of ATP and the ion gradient across the membrane through a rotary catalytic mechanism. Because V-ATPase has structural features similar to those of well-studied F-type ATPase, the structure is expected to highlight the common essence of the torque generation of rotary ATPases. Here, we report a complete model of the extra-membrane domain of the V-ATPase (V1-ATPase) of a thermophilic bacterium, Thermus thermophilus, consisting of three A subunits, three B subunits, one D subunit, and one F subunit. The X-ray structure at 3.9Å resolution provides detailed information about the interactions between A3B3 and DF subcomplexes as well as interactions among the respective subunits, which are defined by the properties of side chains. Asymmetry at the intersubunit interfaces was detected from the structural differences among the three AB pairs in the different reaction states, while the large interdomain motion in the catalytic A subunits was not observed unlike F1 from various species and V1 from Enterococcus hirae. Asymmetry is mainly realized by rigid-body rearrangements of the relative position between A and B subunits. This is consistent with the previous observations by the high-resolution electron microscopy for the whole V-ATPase complexes. Therefore, our result plausibly implies that the essential motion for the torque generation is not the large interdomain movement of the catalytic subunits but the rigid-body rearrangement of subunits.
Organizational Affiliation:
Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan.