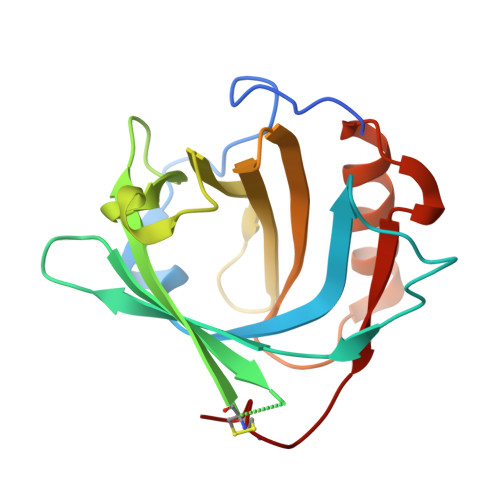
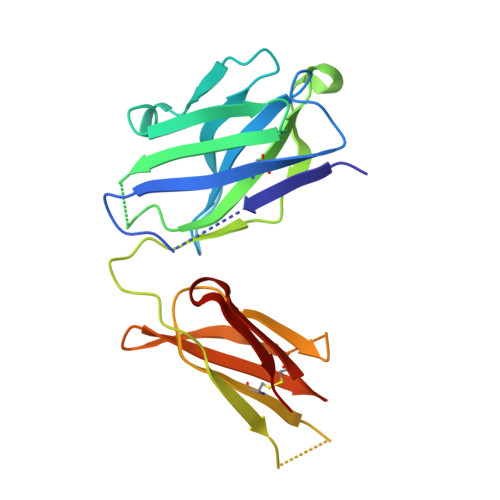
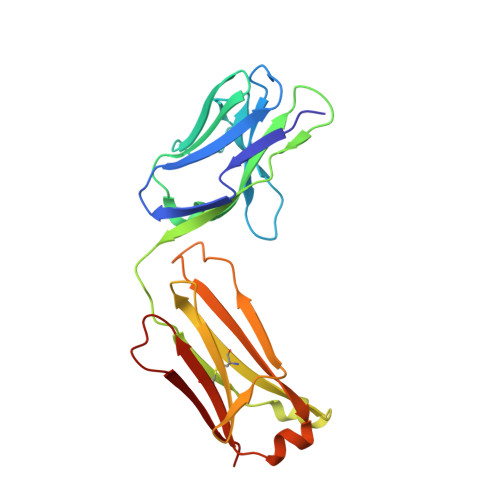
Siderocalin/Lcn2/NGAL/24p3 does not drive apoptosis through gentisic acid mediated iron withdrawal in hematopoietic cell lines.
Correnti, C., Richardson, V., Sia, A.K., Bandaranayake, A.D., Ruiz, M., Suryo Rahmanto, Y., Kovacevic, Z., Clifton, M.C., Holmes, M.A., Kaiser, B.K., Barasch, J., Raymond, K.N., Richardson, D.R., Strong, R.K.(2012) PLoS One 7: e43696-e43696
- PubMed: 22928018
- DOI: https://doi.org/10.1371/journal.pone.0043696
- Primary Citation of Related Structures:
3U0D, 3U9P - PubMed Abstract:
Siderocalin (also lipocalin 2, NGAL or 24p3) binds iron as complexes with specific siderophores, which are low molecular weight, ferric ion-specific chelators. In innate immunity, siderocalin slows the growth of infecting bacteria by sequestering bacterial ferric siderophores. Siderocalin also binds simple catechols, which can serve as siderophores in the damaged urinary tract. Siderocalin has also been proposed to alter cellular iron trafficking, for instance, driving apoptosis through iron efflux via BOCT. An endogenous siderophore composed of gentisic acid (2,5-dihydroxybenzoic acid) substituents was proposed to mediate cellular efflux. However, binding studies reported herein contradict the proposal that gentisic acid forms high-affinity ternary complexes with siderocalin and iron, or that gentisic acid can serve as an endogenous siderophore at neutral pH. We also demonstrate that siderocalin does not induce cellular iron efflux or stimulate apoptosis, questioning the role siderocalin plays in modulating iron metabolism.
Organizational Affiliation:
Division of Basic Sciences, Fred Hutchinson Cancer Research Center, Seattle, Washington, United States of America.