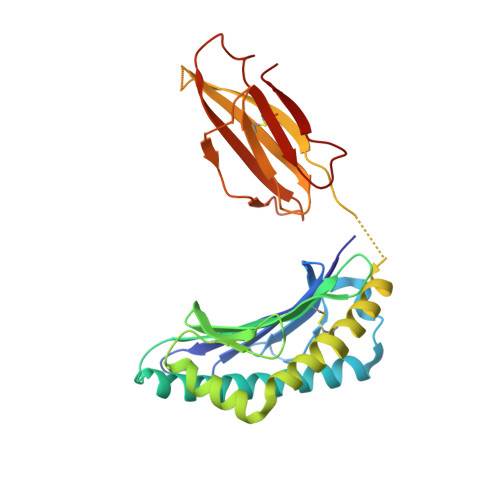
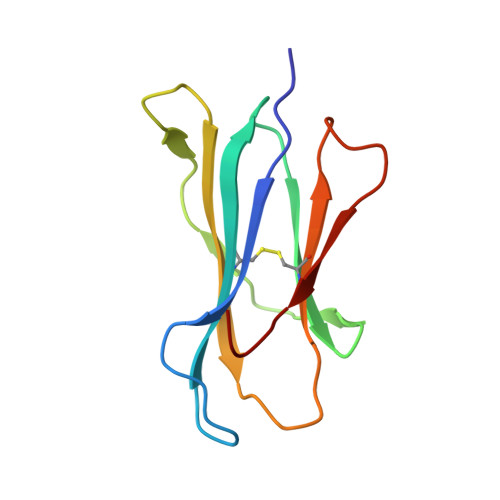
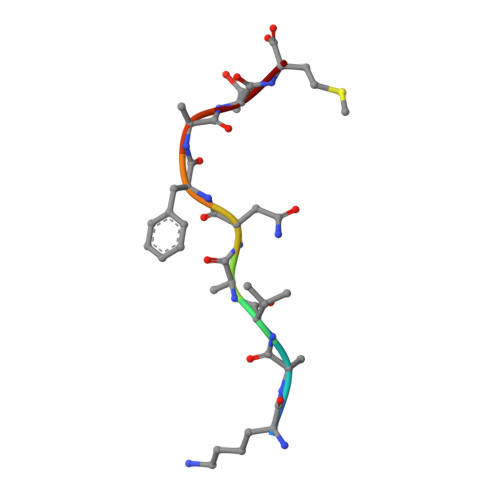
Unexpected T-cell recognition of an altered peptide ligand is driven by reversed thermodynamics.
Allerbring, E.B., Duru, A.D., Uchtenhagen, H., Madhurantakam, C., Tomek, M.B., Grimm, S., Mazumdar, P.A., Friemann, R., Uhlin, M., Sandalova, T., Nygren, P.A., Achour, A.(2012) Eur J Immunol 42: 2990-3000
- PubMed: 22837158
- DOI: https://doi.org/10.1002/eji.201242588
- Primary Citation of Related Structures:
3QUK, 3QUL - PubMed Abstract:
The molecular basis underlying T-cell recognition of MHC molecules presenting altered peptide ligands is still not well-established. A hierarchy of T-cell activation by MHC class I-restricted altered peptide ligands has been defined using the T-cell receptor P14 specific for H-2D(b) in complex with the immunodominant lymphocytic choriomeningitis virus peptide gp33 (KAVYNFATM). While substitution of tyrosine to phenylalanine (Y4F) or serine (Y4S) abolished recognition by P14, the TCR unexpectedly recognized H-2D(b) in complex with the alanine-substituted semiagonist Y4A, which displayed the most significant structural modification. The observed functional hierarchy gp33 > Y4A > Y4S = Y4F was neither due to higher stabilization capacity nor to differences in structural conformation. However, thermodynamic analysis demonstrated that while recognition of the full agonist H-2D(b) /gp33 was strictly enthalpy driven, recognition of the weak agonist H-2D(b) /Y4A was instead entropy driven with a large reduction in the favorable enthalpy term. The fourfold larger negative heat capacity derived for the interaction of P14 with H-2D(b) /gp33 compared with H-2D(b) /Y4A can possibly be explained by higher water entrapment at the TCR/MHC interface, which is also consistent with the measured opposite entropy contributions for the interactions of P14 with both MHCs. In conclusion, this study demonstrates that P14 makes use of different strategies to adapt to structural modifications in the MHC/peptide complex.
Organizational Affiliation:
Center for Infectious Medicine, Department of Medicine, Karolinska University Hospital Huddinge, Karolinska Institutet, Stockholm, Sweden.