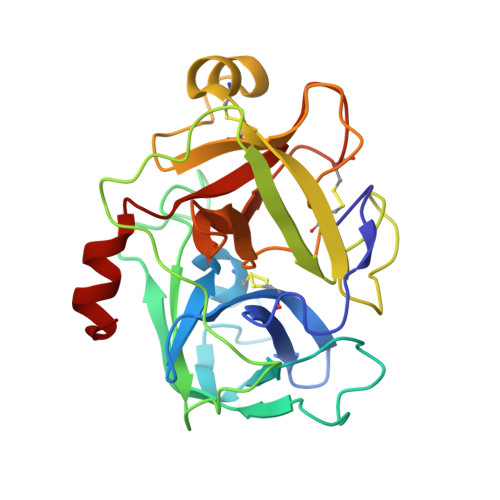
Structure of catalytic domain of Matriptase in complex with Sunflower trypsin inhibitor-1.
Yuan, C., Chen, L., Meehan, E.J., Daly, N., Craik, D.J., Huang, M., Ngo, J.C.(2011) BMC Struct Biol 11: 30-30
- PubMed: 21693064
- DOI: https://doi.org/10.1186/1472-6807-11-30
- Primary Citation of Related Structures:
3P8F, 3P8G - PubMed Abstract:
Matriptase is a type II transmembrane serine protease that is found on the surfaces of epithelial cells and certain cancer cells. Matriptase has been implicated in the degradation of certain extracellular matrix components as well as the activation of various cellular proteins and proteases, including hepatocyte growth factor and urokinase. Sunflower trypsin inhibitor-1 (SFTI-1), a cyclic peptide inhibitor originally isolated from sunflower seeds, exhibits potent inhibitory activity toward matriptase.
Organizational Affiliation:
State Key Lab of Structural Chemistry, Fujian Institute of Research on the Structure of Matter, Chinese Academy of Sciences, Fuzhou, Fujian 350002, China.