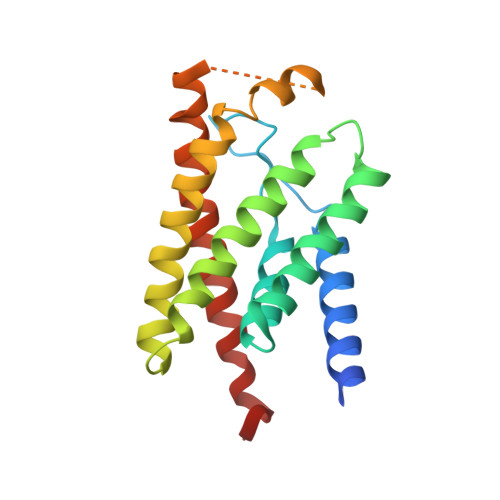
Structure and mechanism of the S component of a bacterial ECF transporter
Zhang, P., Wang, J., Shi, Y.(2010) Nature 468: 717-720
- PubMed: 20972419
- DOI: https://doi.org/10.1038/nature09488
- Primary Citation of Related Structures:
3P5N - PubMed Abstract:
The energy-coupling factor (ECF) transporters, responsible for vitamin uptake in prokaryotes, are a unique family of membrane transporters. Each ECF transporter contains a membrane-embedded, substrate-binding protein (known as the S component), an energy-coupling module that comprises two ATP-binding proteins (known as the A and A' components) and a transmembrane protein (known as the T component). The structure and transport mechanism of the ECF family remain unknown. Here we report the crystal structure of RibU, the S component of the ECF-type riboflavin transporter from Staphylococcus aureus at 3.6-Å resolution. RibU contains six transmembrane segments, adopts a previously unreported transporter fold and contains a riboflavin molecule bound to the L1 loop and the periplasmic portion of transmembrane segments 4-6. Structural analysis reveals the essential ligand-binding residues, identifies the putative transport path and, with sequence alignment, uncovers conserved structural features and suggests potential mechanisms of action among the ECF transporters.
Organizational Affiliation:
Department of Molecular Biology, Lewis Thomas Laboratory, Princeton University, Princeton, New Jersey 08544, USA.