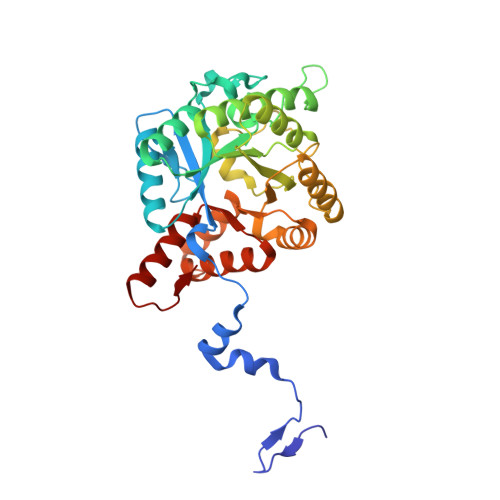
Crystal Structure of Toxoplasma gondii Porphobilinogen Synthase: INSIGHTS ON OCTAMERIC STRUCTURE AND PORPHOBILINOGEN FORMATION.
Jaffe, E.K., Shanmugam, D., Gardberg, A., Dieterich, S., Sankaran, B., Stewart, L.J., Myler, P.J., Roos, D.S.(2011) J Biol Chem 286: 15298-15307
- PubMed: 21383008
- DOI: https://doi.org/10.1074/jbc.M111.226225
- Primary Citation of Related Structures:
3OBK - PubMed Abstract:
Porphobilinogen synthase (PBGS) is essential for heme biosynthesis, but the enzyme of the protozoan parasite Toxoplasma gondii (TgPBGS) differs from that of its human host in several important respects, including subcellular localization, metal ion dependence, and quaternary structural dynamics. We have solved the crystal structure of TgPBGS, which contains an octamer in the crystallographic asymmetric unit. Crystallized in the presence of substrate, each active site contains one molecule of the product porphobilinogen. Unlike prior structures containing a substrate-derived heterocycle directly bound to an active site zinc ion, the product-bound TgPBGS active site contains neither zinc nor magnesium, placing in question the common notion that all PBGS enzymes require an active site metal ion. Unlike human PBGS, the TgPBGS octamer contains magnesium ions at the intersections between pro-octamer dimers, which are presumed to function in allosteric regulation. TgPBGS includes N- and C-terminal regions that differ considerably from previously solved crystal structures. In particular, the C-terminal extension found in all apicomplexan PBGS enzymes forms an intersubunit β-sheet, stabilizing a pro-octamer dimer and preventing formation of hexamers that can form in human PBGS. The TgPBGS structure suggests strategies for the development of parasite-selective PBGS inhibitors.
Organizational Affiliation:
Institute for Cancer Research, Fox Chase Cancer Center, Philadelphia, Pennsylvania 19111, USA. eileen.jaffe@fccc.edu