Investigation of 2-Fold Disorder of Inhibitors and Relative Potency by Crystallizations of HIV-1 Protease in Ritonavir and Saquinavir Mixtures
Olajuyigbe, F.M., Demitri, N., Geremia, S.(2011) Cryst Growth Des 11: 4378-4385
Experimental Data Snapshot
(2011) Cryst Growth Des 11: 4378-4385
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Protease | 100 | Human immunodeficiency virus 1 | Mutation(s): 5 Gene Names: pol EC: 3.4.23.16 | 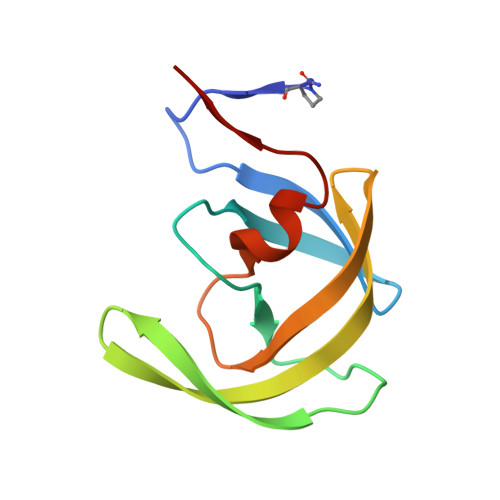 | |
UniProt | |||||
Find proteins for P03367 (Human immunodeficiency virus type 1 group M subtype B (isolate BRU/LAI)) Explore P03367 Go to UniProtKB: P03367 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P03367 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ROC Query on ROC | E [auth A], Z [auth D] | (2S)-N-[(2S,3R)-4-[(2S,3S,4aS,8aS)-3-(tert-butylcarbamoyl)-3,4,4a,5,6,7,8,8a-octahydro-1H-isoquinolin-2-yl]-3-hydroxy-1
-phenyl-butan-2-yl]-2-(quinolin-2-ylcarbonylamino)butanediamide C38 H50 N6 O5 QWAXKHKRTORLEM-UGJKXSETSA-N |  | ||
| GOL Query on GOL | I [auth A], N [auth B], Y [auth C] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| DMS Query on DMS | AA [auth D], F [auth A], M [auth B], T [auth C], U [auth C] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | G [auth A], K [auth A], R [auth B], X [auth C] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| CL Query on CL | H [auth A] L [auth A] O [auth B] P [auth B] V [auth C] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| NA Query on NA | J [auth A], Q [auth B], S [auth B] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Binding Affinity Annotations | |||
|---|---|---|---|
| ID | Source | Binding Affinity | |
| ROC | BindingDB: 3NDU | Ki: min: 0.04, max: 138 (nM) from 29 assay(s) | |
| Kd: min: 0.31, max: 67 (nM) from 6 assay(s) | |||
| IC50: min: 0.2, max: 270 (nM) from 16 assay(s) | |||
| -TΔS: min: -8.03e+1, max: -2.80e+1 (kJ/mol) from 47 assay(s) | |||
| ΔH: min: -3.18e+1, max: 43.47 (kJ/mol) from 47 assay(s) | |||
| ΔG: min: -6.23e+1, max: -3.55e+1 (kJ/mol) from 44 assay(s) | |||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_000454 (ROC) Query on PRD_000454 | E [auth A], Z [auth D] | Saquinavir | Peptide-like / Inhibitor |  | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 51.19 | α = 90 |
| b = 62.705 | β = 98.32 |
| c = 59.296 | γ = 90 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data collection |
| SHELX | model building |
| SHELXL-97 | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| SHELX | phasing |