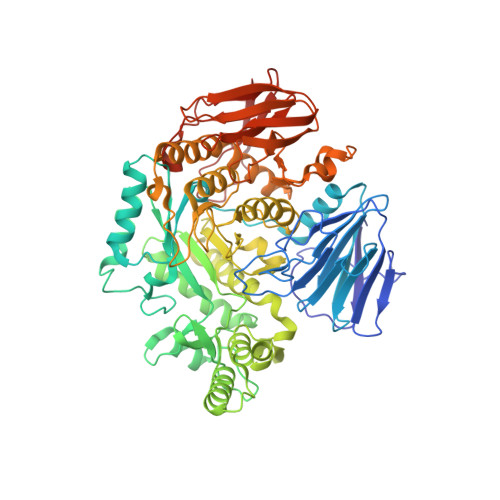
The Crystal Structures Of The Glycoside Hydrolase (Family 31) From Ruminococcus Obeum Atcc 29174
Tan, K., Tesar, C., Wilton, R., Keigher, L., Babnigg, G., Joachimiak, A.(2010) FASEB J 24: 3939-3949
- PubMed: 20581222
- DOI: https://doi.org/10.1096/fj.10-156257
- Primary Citation of Related Structures:
3M46, 3M6D, 3MKK, 3N04 - PubMed Abstract:
The human intestine harbors a large number of microbes forming a complex microbial community that greatly affects the physiology and pathology of the host. In the human gut microbiome, the enrichment in certain protein gene families appears to be widespread. They include enzymes involved in carbohydrate metabolism such as glucoside hydrolases of dietary polysaccharides and glycoconjugates. We report the crystal structures (wild type, 2 mutants, and a mutant/substrate complex) and the enzymatic activity of a recombinant α-glucosidase from human gut bacterium Ruminococcus obeum. The first ever protein structures from this bacterium reveal a structural homologue to human intestinal maltase-glucoamylase with a highly conserved catalytic domain and reduced auxiliary domains. The α-glucosidase, a member of GH31 family, shows substrate preference for α(1-6) over α(1-4) glycosidic linkages and produces glucose from isomaltose as well as maltose. The preference can be switched by a single mutation at its active site, suggestive of widespread adaptation to utilization of a variety of polysaccharides by intestinal micro-organisms as energy resources.
Organizational Affiliation:
Argonne National Laboratory, 9700 South Cass Avenue, Argonne, IL 60439, USA.