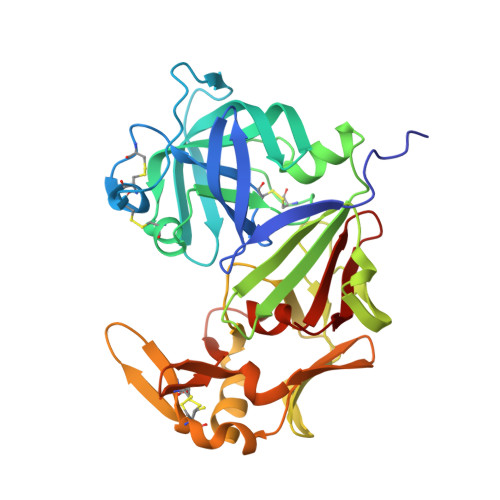
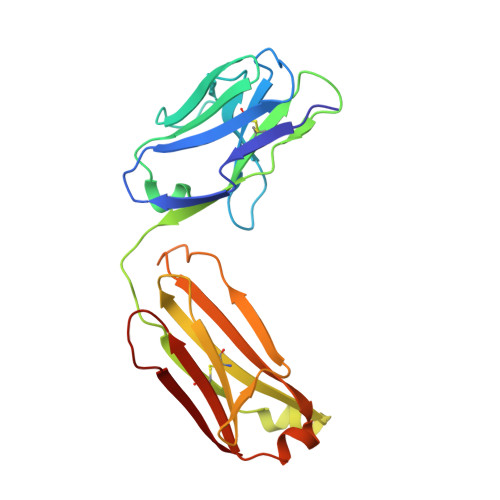
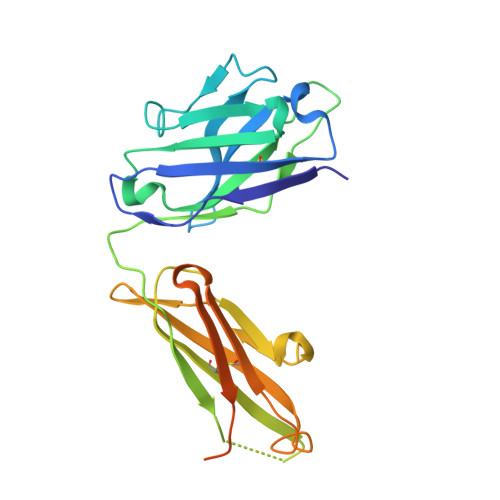
Mechanisms of allergen-antibody interaction of cockroach allergen Bla g 2 with monoclonal antibodies that inhibit IgE antibody binding.
Glesner, J., Wunschmann, S., Li, M., Gustchina, A., Wlodawer, A., Himly, M., Chapman, M.D., Pomes, A.(2011) PLoS One 6: e22223-e22223
- PubMed: 21789239
- DOI: https://doi.org/10.1371/journal.pone.0022223
- Primary Citation of Related Structures:
3LIZ - PubMed Abstract:
Cockroach allergy is strongly associated with asthma, and involves the production of IgE antibodies against inhaled allergens. Reports of conformational epitopes on inhaled allergens are limited. The conformational epitopes for two specific monoclonal antibodies (mAb) that interfere with IgE antibody binding were identified by X-ray crystallography on opposite sites of the quasi-symmetrical cockroach allergen Bla g 2.
Organizational Affiliation:
INDOOR Biotechnologies, Inc, Charlottesville, Virginia, United States of America.