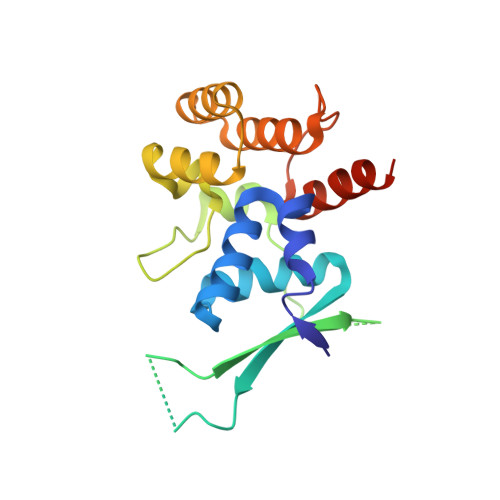
Insights into eukaryotic DNA priming from the structure and functional interactions of the 4Fe-4S cluster domain of human DNA primase.
Vaithiyalingam, S., Warren, E.M., Eichman, B.F., Chazin, W.J.(2010) Proc Natl Acad Sci U S A 107: 13684-13689
- PubMed: 20643958
- DOI: https://doi.org/10.1073/pnas.1002009107
- Primary Citation of Related Structures:
3L9Q - PubMed Abstract:
DNA replication requires priming of DNA templates by enzymes known as primases. Although DNA primase structures are available from archaea and bacteria, the mechanism of DNA priming in higher eukaryotes remains poorly understood in large part due to the absence of the structure of the unique, highly conserved C-terminal regulatory domain of the large subunit (p58C). Here, we present the structure of this domain determined to 1.7-A resolution by X-ray crystallography. The p58C structure reveals a novel arrangement of an evolutionarily conserved 4Fe-4S cluster buried deeply within the protein core and is not similar to any known protein structure. Analysis of the binding of DNA to p58C by fluorescence anisotropy measurements revealed a strong preference for ss/dsDNA junction substrates. This approach was combined with site-directed mutagenesis to confirm that the binding of DNA occurs to a distinctively basic surface on p58C. A specific interaction of p58C with the C-terminal domain of the intermediate subunit of replication protein A (RPA32C) was identified and characterized by isothermal titration calorimetry and NMR. Restraints from NMR experiments were used to drive computational docking of the two domains and generate a model of the p58C-RPA32C complex. Together, our results explain functional defects in human DNA primase mutants and provide insights into primosome loading on RPA-coated ssDNA and regulation of primase activity.
Organizational Affiliation:
Department of Biochemistry, Center for Structural Biology, Vanderbilt University, Nashville, TN 37232-8725, USA.