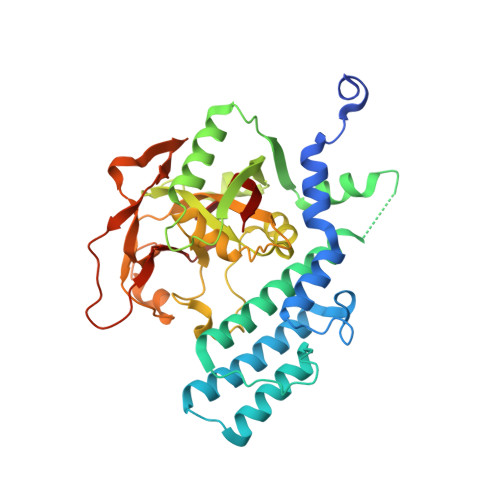
Crystal structure of the catalytic domain of human PARP2 in complex with PARP inhibitor ABT-888.
Karlberg, T., Hammarstrom, M., Schutz, P., Svensson, L., Schuler, H.(2010) Biochemistry 49: 1056-1058
- PubMed: 20092359
- DOI: https://doi.org/10.1021/bi902079y
- Primary Citation of Related Structures:
3KCZ, 3KJD - PubMed Abstract:
Poly-ADP-ribose polymerases (PARPs) catalyze transfer of ADP-ribose from NAD(+) to specific residues in their substrate proteins or to growing ADP-ribose chains. PARP activity is involved in processes such as chromatin remodeling, transcription control, and DNA repair. Inhibitors of PARP activity may be useful in cancer therapy. PARP2 is the family member that is most similar to PARP1, and the two can act together as heterodimers. We used X-ray crystallography to determine two structures of the catalytic domain of human PARP2: the complexes with PARP inhibitors 3-aminobenzamide and ABT-888. These results contribute to our understanding of structural features and compound properties that can be employed to develop selective inhibitors of human ADP-ribosyltransferases.
Organizational Affiliation:
Structural Genomics Consortium, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, Scheeles vag 2, 17177 Stockholm, Sweden.