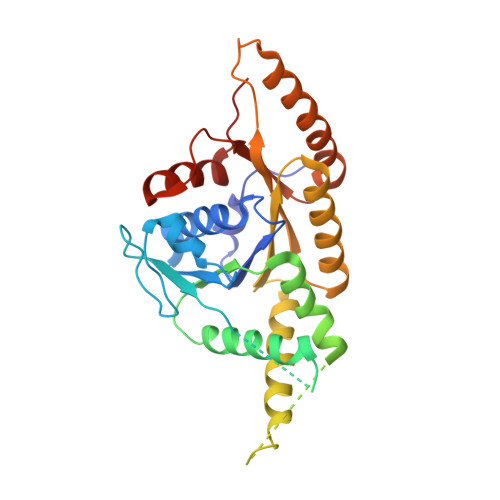
Structural insights into tail-anchored protein binding and membrane insertion by Get3.
Bozkurt, G., Stjepanovic, G., Vilardi, F., Amlacher, S., Wild, K., Bange, G., Favaloro, V., Rippe, K., Hurt, E., Dobberstein, B., Sinning, I.(2009) Proc Natl Acad Sci U S A 106: 21131-21136
- PubMed: 19948960
- DOI: https://doi.org/10.1073/pnas.0910223106
- Primary Citation of Related Structures:
3IQW, 3IQX - PubMed Abstract:
Tail-anchored (TA) membrane proteins are involved in a variety of important cellular functions, including membrane fusion, protein translocation, and apoptosis. The ATPase Get3 (Asna1, TRC40) was identified recently as the endoplasmic reticulum targeting factor of TA proteins. Get3 consists of an ATPase and alpha-helical subdomain enriched in methionine and glycine residues. We present structural and biochemical analyses of Get3 alone as well as in complex with a TA protein, ribosome-associated membrane protein 4 (Ramp4). The ATPase domains form an extensive dimer interface that encloses 2 nucleotides in a head-to-head orientation and a zinc ion. Amide proton exchange mass spectrometry shows that the alpha-helical subdomain of Get3 displays considerable flexibility in solution and maps the TA protein-binding site to the alpha-helical subdomain. The non-hydrolyzable ATP analogue AMPPNP-Mg(2+)- and ADP-Mg(2+)-bound crystal structures representing the pre- and posthydrolysis states are both in a closed form. In the absence of a TA protein cargo, ATP hydrolysis does not seem to be possible. Comparison with the ADP.AlF(4)(-)-bound structure representing the transition state (Mateja A, et al. (2009) Nature 461:361-366) indicates how the presence of a TA protein is communicated to the ATP-binding site. In vitro membrane insertion studies show that recombinant Get3 inserts Ramp4 in a nucleotide- and receptor-dependent manner. Although ATP hydrolysis is not required for Ramp4 insertion per se, it seems to be required for efficient insertion. We postulate that ATP hydrolysis is needed to release Get3 from its receptor. Taken together, our results provide mechanistic insights into posttranslational targeting of TA membrane proteins by Get3.
Organizational Affiliation:
Heidelberg University Biochemistry Center, Im Neuenheimer Feld 328, D-69120 Heidelberg, Germany.