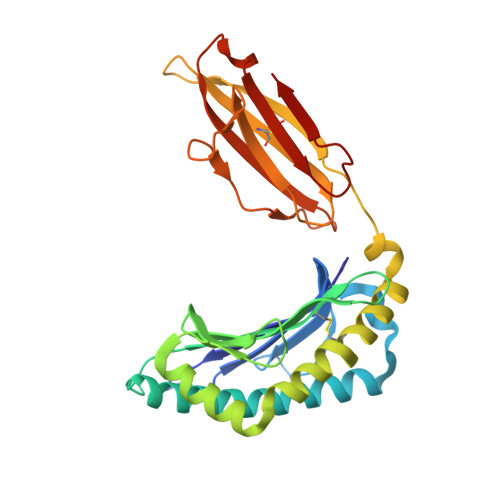
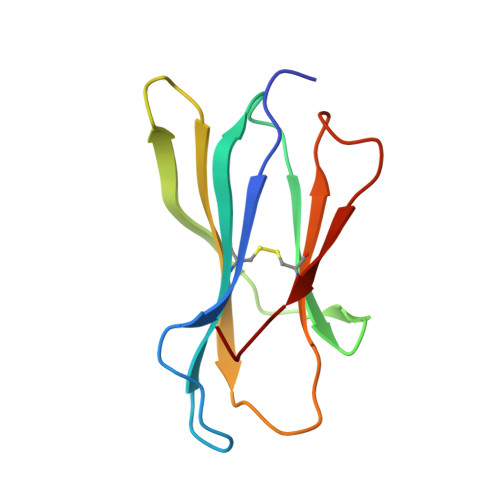
Structural basis of the CD8alphabeta/MHC class i interaction: focused recognition orients CD8beta to a T cell proximal position
Wang, R., Natarajan, K., Margulies, D.H.(2009) J Immunol 183: 2554-2564
- PubMed: 19625641
- DOI: https://doi.org/10.4049/jimmunol.0901276
- Primary Citation of Related Structures:
3DMM, 3ECB - PubMed Abstract:
In the immune system, B cells, dendritic cells, NK cells, and T lymphocytes all respond to signals received via ligand binding to receptors and coreceptors. Although the specificity of T cell recognition is determined by the interaction of T cell receptors with MHC/peptide complexes, the development of T cells in the thymus and their sensitivity to Ag are also dependent on coreceptor molecules CD8 (for MHC class I (MHCI)) and CD4 (for MHCII). The CD8alphabeta heterodimer is a potent coreceptor for T cell activation, but efforts to understand its function fully have been hampered by ignorance of the structural details of its interactions with MHCI. In this study we describe the structure of CD8alphabeta in complex with the murine MHCI molecule H-2D(d) at 2.6 A resolution. The focus of the CD8alphabeta interaction is the acidic loop (residues 222-228) of the alpha3 domain of H-2D(d). The beta subunit occupies a T cell membrane proximal position, defining the relative positions of the CD8alpha and CD8beta subunits. Unlike the CD8alphaalpha homodimer, CD8alphabeta does not contact the MHCI alpha(2)- or beta(2)-microglobulin domains. Movements of the CD8alpha CDR2 and CD8beta CDR1 and CDR2 loops as well as the flexibility of the H-2D(d) CD loop facilitate the monovalent interaction. The structure resolves inconclusive data on the topology of the CD8alphabeta/MHCI interaction, indicates that CD8beta is crucial in orienting the CD8alphabeta heterodimer, provides a framework for understanding the mechanistic role of CD8alphabeta in lymphoid cell signaling, and offers a tangible context for design of structurally altered coreceptors for tumor and viral immunotherapy.
Organizational Affiliation:
Molecular Biology Section, Laboratory of Immunology, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.