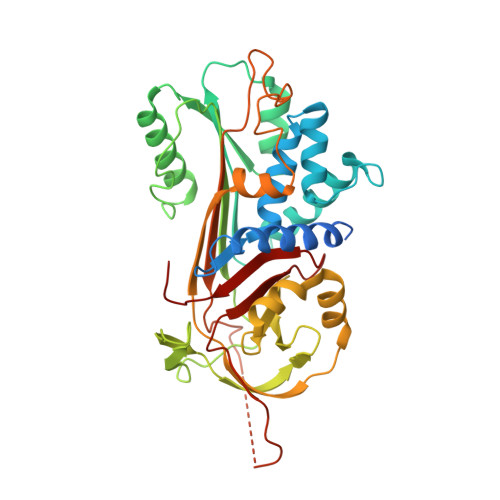
Crystallographic and cellular characterisation of two mechanisms stabilising the native fold of alpha1-antitrypsin: implications for disease and drug design.
Gooptu, B., Miranda, E., Nobeli, I., Mallya, M., Purkiss, A., Brown, S.C., Summers, C., Phillips, R.L., Lomas, D.A., Barrett, T.E.(2009) J Mol Biol 387: 857-868
- PubMed: 19232354
- DOI: https://doi.org/10.1016/j.jmb.2009.01.069
- Primary Citation of Related Structures:
3DRM, 3DRU - PubMed Abstract:
The common Z mutant (Glu342Lys) of alpha(1)-antitrypsin results in the formation of polymers that are retained within hepatocytes. This causes liver disease whilst the plasma deficiency of an important proteinase inhibitor predisposes to emphysema. The Thr114Phe and Gly117Phe mutations border a surface cavity identified as a target for rational drug design. These mutations preserve inhibitory activity but reduce the polymerisation of wild-type native alpha(1)-antitrypsin in vitro and increase secretion in a Xenopus oocyte model of disease. To understand these effects, we have crystallised both mutants and solved their structures. The 2.2 A structure of Thr114Phe alpha(1)-antitrypsin demonstrates that the effects of the mutation are mediated entirely by well-defined partial cavity blockade and allows in silico screening of fragments capable of mimicking the effects of the mutation. The Gly117Phe mutation operates differently, repacking aromatic side chains in the helix F-beta-sheet A interface to induce a half-turn downward shift of the adjacent F helix. We have further characterised the effects of these two mutations in combination with the Z mutation in a eukaryotic cell model of disease. Both mutations increase the secretion of Z alpha(1)-antitrypsin in the native conformation, but the double mutants remain more polymerogenic than the wild-type (M) protein. Taken together, these data support different mechanisms by which the Thr114Phe and Gly117Phe mutations stabilise the native fold of alpha(1)-antitrypsin and increase secretion of monomeric protein in cell models of disease.
Organizational Affiliation:
School of Crystallography, Birkbeck College, University of London, London, UK. b.gooptu@mail.cryst.bbk.ac.uk