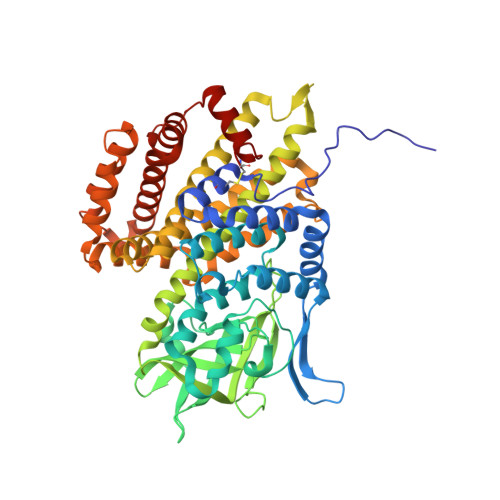
Structure and DNA binding of alkylation response protein AidB.
Bowles, T., Metz, A.H., O'Quin, J., Wawrzak, Z., Eichman, B.F.(2008) Proc Natl Acad Sci U S A 105: 15299-15304
- PubMed: 18829440
- DOI: https://doi.org/10.1073/pnas.0806521105
- Primary Citation of Related Structures:
3DJL - PubMed Abstract:
Exposure of Escherichia coli to alkylating agents activates expression of AidB in addition to DNA repair proteins Ada, AlkA, and AlkB. AidB was recently shown to possess a flavin adenine dinucleotide (FAD) cofactor and to bind to dsDNA, implicating it as a flavin-dependent DNA repair enzyme. However, the molecular mechanism by which AidB acts to reduce the mutagenic effects of specific DNA alkylators is unknown. We present a 1.7-A crystal structure of AidB, which bears superficial resemblance to the acyl-CoA dehydrogenase superfamily of flavoproteins. The structure reveals a unique quaternary organization and a distinctive FAD active site that provides a rationale for AidB's limited dehydrogenase activity. A highly electropositive C-terminal domain not present in structural homologs was identified by mutational analysis as the DNA binding site. Structural analysis of the DNA and FAD binding sites provides evidence against AidB-catalyzed DNA repair and supports a model in which AidB acts to prevent alkylation damage by protecting DNA and destroying alkylating agents that have yet to reach their DNA target.
Organizational Affiliation:
Department of Biological Sciences and Center for Structural Biology, Vanderbilt University, Nashville, TN 37232, USA.