How a T cell receptor-like antibody recognizes major histocompatibility complex-bound peptide
Mareeva, T., Martinez-Hackert, E., Sykulev, Y.(2008) J Biol Chem 283: 29053-29059
- PubMed: 18703505
- DOI: https://doi.org/10.1074/jbc.M804996200
- Primary Citation of Related Structures:
3CVH, 3CVI - PubMed Abstract:
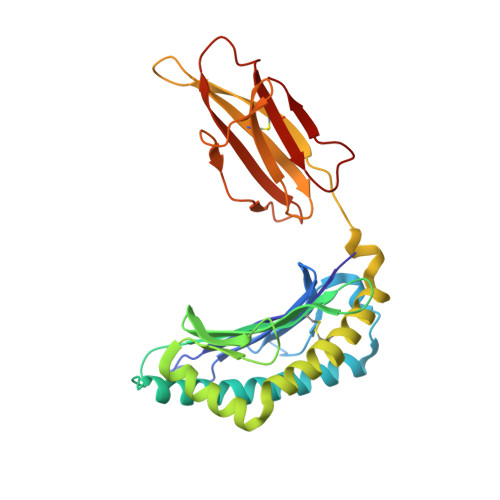
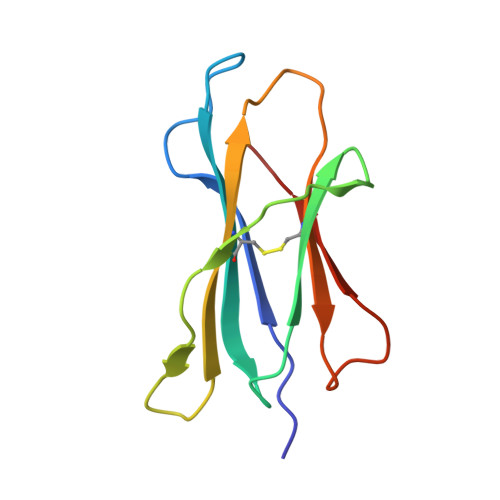
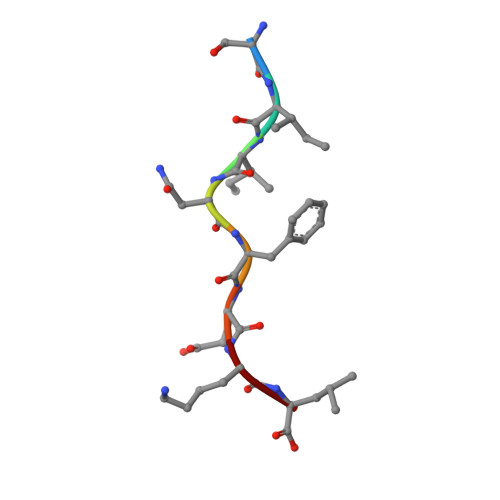
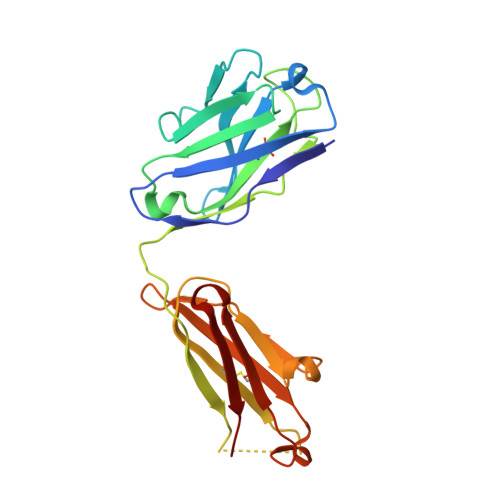
We determined the crystal structures of the T cell receptor (TCR)-like antibody 25-D1.16 Fab fragment bound to a complex of SIINFEKL peptide from ovalbumin and the H-2K(b) molecule. Remarkably, this antibody directly "reads" the structure of the major histocompatibility complex (MHC)-bound peptide, employing the canonical diagonal binding mode utilized by most TCRs. This is in marked contrast with another TCR-like antibody, Hyb3, bound to melanoma peptide MAGE-A1 in association with HLA-A1 MHC class I. Hyb3 assumes a non-canonical orientation over its cognate peptide-MHC and appears to recognize a conformational epitope in which the MHC contribution is dominant. We conclude that TCR-like antibodies can recognize MHC-bound peptide via two different mechanisms: one is similar to that exploited by the preponderance of TCRs and the other requires a non-canonical antibody orientation over the peptide-MHC complex.
Organizational Affiliation:
Department of Microbiology and Immunology and the Kimmel Cancer Center, Thomas Jefferson University, Philadelphia, Pennsylvania 19107, USA.