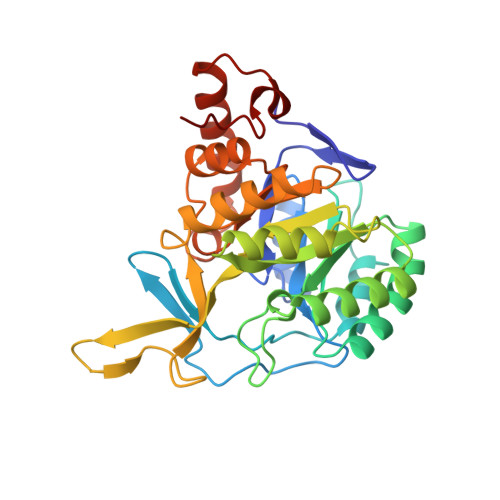
Crystal structure of Trypanosoma cruzi dihydroorotate dehydrogenase from Y strain
Pinheiro, M.P., Iulek, J., Cristina Nonato, M.(2008) Biochem Biophys Res Commun 369: 812-817
- PubMed: 18302934
- DOI: https://doi.org/10.1016/j.bbrc.2008.02.074
- Primary Citation of Related Structures:
3C3N - PubMed Abstract:
Trypanosoma cruzi is the etiological agent of Chagas' disease, a pathogenesis that affects millions of people in Latin America. Here, we report the crystal structure of dihydroorotate dehydrogenase (DHODH) from T. cruzi strain Y solved at 2.2A resolution. DHODH is a flavin mononucleotide containing enzyme, which catalyses the oxidation of l-dihydroorotate to orotate, the fourth step and only redox reaction in the de novo biosynthesis of pyrimidine nucleotides. Genetic studies have shown that DHODH is essential for T. cruzi survival, validating the idea that this enzyme can be considered an attractive target for the development of antichagasic drugs. In our work, a detailed analysis of T. cruzi DHODH crystal structure has allowed us to suggest potential sites to be further exploited for the design of highly specific inhibitors through the technology of structure-based drug design.
Organizational Affiliation:
Laboratório de Cristalografia de Proteínas, Departamento de Física e Química, Faculdade de Ciências Farmacêuticas de Ribeirão Preto, Universidade de São Paulo, Ribeirão Preto S.P. 14040-903, Brazil.