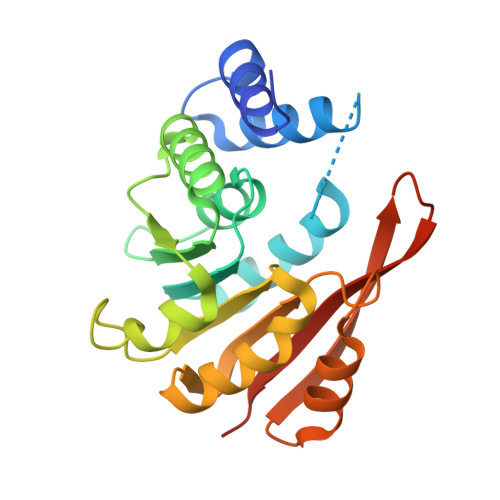
Crystal structures of the Apo and Holo form of rat catechol-O-methyltransferase
Tsuji, E., Okazaki, K., Isaji, M., Takeda, K.(2009) J Struct Biol 165: 133-139
- PubMed: 19111934
- DOI: https://doi.org/10.1016/j.jsb.2008.11.012
- Primary Citation of Related Structures:
2ZLB, 2ZTH - PubMed Abstract:
Catechol-O-methyltransferase (COMT, EC 2.1.1.6) is a monomeric enzyme that catalyzes the transfer of a methyl group from S-adenosyl-l-methionine (AdoMet) to the phenolic oxygen of substituted catechols. Although the inhibitor recognition pattern and AdoMet site have already been studied crystallographically, structural information on the catalytic cycle of COMT has not yet been obtained. In this study, comparison of the co-factor and inhibitor-bound structures revealed that the Apo form of COMT shows a conformational change and there was no cleft corresponding to the AdoMet-binding site; the overall structure was partially open form and the substrate recognition site was not clearly defined. The Holo form of COMT was similar to the quaternary structure except for the beta6-beta7 and alpha2-alpha3 ligand recognition loops. These conformational changes provide a deeper insight into the structural events occurring in reactions catalyzed by AdoMet.
Organizational Affiliation:
Molecular Design Research, R&D Kissei Pharmaceutical Co., Ltd. 4365-1 Kashiwabara, Hotaka, Azumino-city, Nagano 399-8304, Japan. eiichi_tsuji@pharm.kissei.co.jp