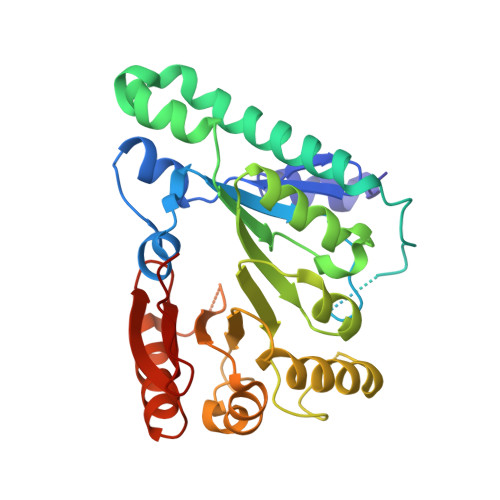
Crystal Structure of the Radical SAM Enzyme Catalyzing Tricyclic Modified Base Formation in tRNA
Suzuki, Y., Noma, A., Suzuki, T., Senda, M., Senda, T., Ishitani, R., Nureki, O.(2007) J Mol Biol 372: 1204-1214
- PubMed: 17727881
- DOI: https://doi.org/10.1016/j.jmb.2007.07.024
- Primary Citation of Related Structures:
2Z2U - PubMed Abstract:
Wyosine and its derivatives, such as wybutosine, found in eukaryotic and archaeal tRNAs, are tricyclic hypermodified nucleosides. In eukaryotes, wybutosine exists exclusively in position 37, 3'-adjacent to the anticodon, of tRNA(Phe), where it ensures correct translation by stabilizing the codon-anticodon base-pairing during the ribosomal decoding process. Recent studies revealed that the wyosine biosynthetic pathway consists of multistep enzymatic reactions starting from a guanosine residue. Among these steps, TYW1 catalyzes the second step to form the tricyclic ring structure, by cyclizing N(1)-methylguanosine. In this study, we solved the crystal structure of TYW1 from Methanocaldococcus jannaschii at 2.4 A resolution. TYW1 assumes an incomplete TIM barrel with (alpha/beta)(6) topology, which closely resembles the reported structures of radical SAM enzymes. Hence, TYW1 was considered to catalyze the cyclization reaction by utilizing the radical intermediate. Comparison with other radical SAM enzymes allowed us to build a model structure complexed with S-adenosylmethionine and two [4Fe-4S] clusters. Mutational analyses in yeast supported the validity of this complex model structure, which provides a structural insight into the radical reaction involving two [4Fe-4S] clusters to create a complex tricyclic base.
Organizational Affiliation:
Department of Biological Information, Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, B34 4259 Nagatsuta-cho, Midori-ku, Yokohama-shi, Kanagawa 226-8501, Japan.