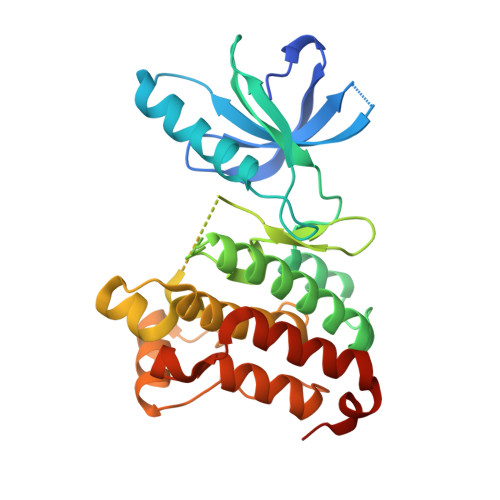
Stability and Solubility Engineering of the Ephb4 Tyrosine Kinase Catalytic Domain Using a Rationally Designed Synthetic Library.
Overman, R.C., Green, I., Truman, C.M., Read, J.A., Embrey, K.J., Mcalister, M.S.B., Attwood, T.K.(2013) Protein Eng Des Sel 26: 695
- PubMed: 23840071
- DOI: https://doi.org/10.1093/protein/gzt032
- Primary Citation of Related Structures:
2YN8 - PubMed Abstract:
The inability to generate soluble, correctly folded recombinant protein is often a barrier to successful structural and functional studies. Access to affordable synthetic genes has, however, made it possible to design, make and test many more variants of a target protein to identify suitable constructs. We have used rational design and gene synthesis to create a controlled randomised library of the EphB4 receptor tyrosine kinase, with the aim of obtaining soluble, purifiable and active catalytic domain material at multi-milligram levels in Escherichia coli. Three main parameters were tested in designing the library--construct length, functional mutations and stability grafting. These variables were combined to generate a total of 9720 possible variants. The screening of 480 clones generated a 3% hit rate, with a purifiable solubility of up to 15 mg/L for some EphB4 constructs that was largely independent of construct length. Sequencing of the positive clones revealed a pair of hydrophobic core mutations that were key to obtaining soluble material. A minimal kinase domain construct containing these two mutations exhibited a +4.5°C increase in thermal stability over the wild-type protein. These approaches will be broadly applicable for solubility engineering of many different protein target classes. Atomic coordinates and structural factors have been deposited in PDB under the accession 2yn8 (EphB4 HP + staurosporine).
Organizational Affiliation:
Discovery Sciences, AstraZeneca PLC, Alderley Park, Cheshire SK10 4TG, UK.