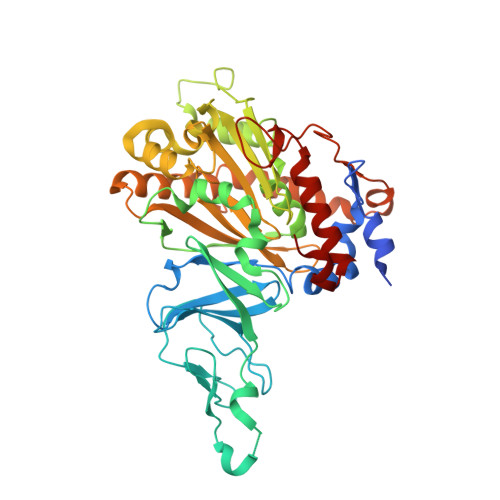
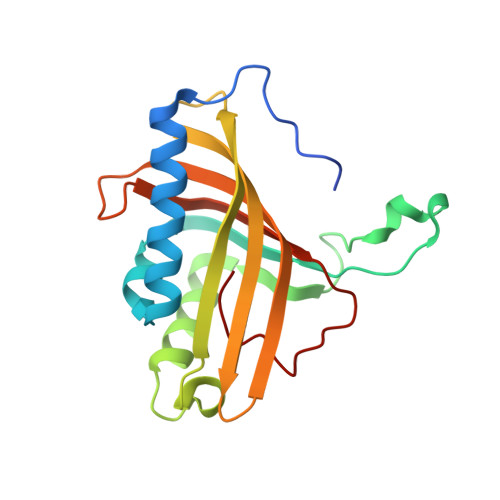
Structural Insights Into the Metabolism of 2-Chlorodibenzofuran by an Evolved Biphenyl Dioxygenase.
Kumar, P., Mohammadi, M., Dhindwal, S., Pham, T.T., Bolin, J.T., Sylvestre, M.(2012) Biochem Biophys Res Commun 421: 757
- PubMed: 22546558
- DOI: https://doi.org/10.1016/j.bbrc.2012.04.078
- Primary Citation of Related Structures:
2YFL - PubMed Abstract:
The biphenyl dioxygenase of Burkholderia xenovorans LB400 (BphAE(LB400)) is a Rieske-type oxygenase that catalyzes the stereospecific oxygenation of many heterocyclic aromatics including dibenzofuran. In a previous work, we evolved BphAE(LB400) and obtained BphAE(RR41). This variant metabolizes dibenzofuran and 2-chlorodibenzofuran more efficiently than BphAE(LB400). However, the regiospecificity of BphAE(RR41) toward these substrates differs. Dibenzofuran is metabolized principally through a lateral dioxygenation whereas 2-chlorodibenzofuran is metabolized principally through an angular dioxygenation. In order to explain this difference, we examined the crystal structures of both substrate-bound forms of BphAE(RR41) obtained under anaerobic conditions. This structure analysis, in combination with biochemical data for a Ser283Gly mutant provided evidences that the substrate is compelled to move after oxygen-binding in BphAE(RR41):dibenzofuran. In BphAE(RR41):2-chlorodibenzofuran, the chlorine atom is close to the side chain of Ser283. This contact is missing in the BphAE(RR41):dibenzofuran, and strong enough in the BphAE(RR41):2-chlorodibenzofuran to help prevent substrate movement during the catalytic reaction.
Organizational Affiliation:
Department of Biological Sciences and Center for Cancer Research, Purdue University, West Lafayette, IN 47907, USA.