Remote Control of Regioselectivity in Acyl-Acyl Carrier Protein-Desaturases.
Guy, J.E., Whittle, E., Moche, M., Lengqvist, J., Lindqvist, Y., Shanklin, J.(2011) Proc Natl Acad Sci U S A 108: 16594
- PubMed: 21930947
- DOI: https://doi.org/10.1073/pnas.1110221108
- Primary Citation of Related Structures:
2XZ0, 2XZ1 - PubMed Abstract:
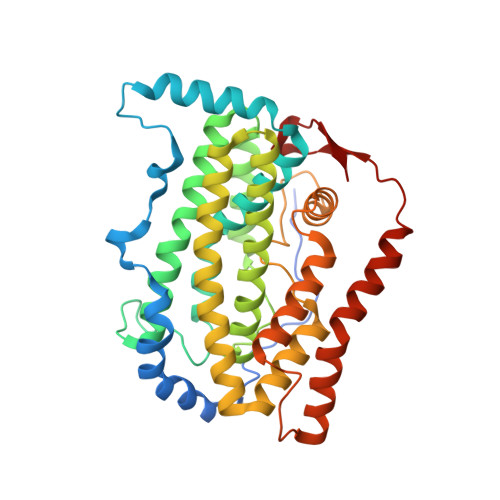
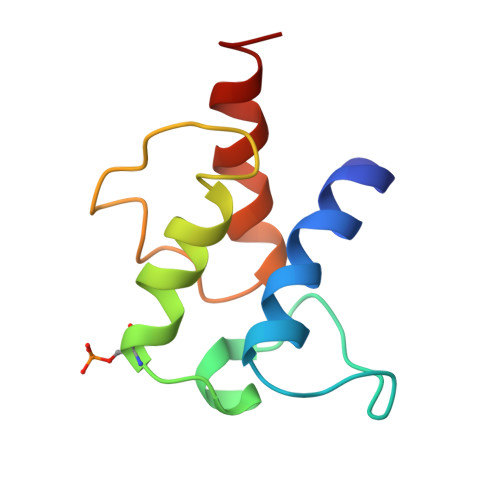
Regiospecific desaturation of long-chain saturated fatty acids has been described as approaching the limits of the discriminatory power of enzymes because the substrate entirely lacks distinguishing features close to the site of dehydrogenation. To identify the elusive mechanism underlying regioselectivity, we have determined two crystal structures of the archetypal Δ9 desaturase from castor in complex with acyl carrier protein (ACP), which show the bound ACP ideally situated to position C9 and C10 of the acyl chain adjacent to the diiron active site for Δ9 desaturation. Analysis of the structures and modeling of the complex between the highly homologous ivy Δ4 desaturase and ACP, identified a residue located at the entrance to the binding cavity, Asp280 in the castor desaturase (Lys275 in the ivy desaturase), which is strictly conserved within Δ9 and Δ4 enzymes but differs between them. We hypothesized that interaction between Lys275 and the phosphate of the pantetheine, seen in the ivy model, is key to positioning C4 and C5 adjacent to the diiron center for Δ4 desaturation. Mutating castor Asp280 to Lys resulted in a major shift from Δ9 to Δ4 desaturation. Thus, interaction between desaturase side-chain 280 and phospho-serine 38 of ACP, approximately 27 Å from the site of double-bond formation, predisposes ACP binding that favors either Δ9 or Δ4 desaturation via repulsion (acidic side chain) or attraction (positively charged side chain), respectively. Understanding the mechanism underlying remote control of regioselectivity provides the foundation for reengineering desaturase enzymes to create designer chemical feedstocks that would provide alternatives to those currently obtained from petrochemicals.
Organizational Affiliation:
Department of Medical Biochemistry and Biophysics, Molecular Structural Biology, Karolinska Institutet, Tomtebodavägen 6, S-171 77 Stockholm, Sweden.