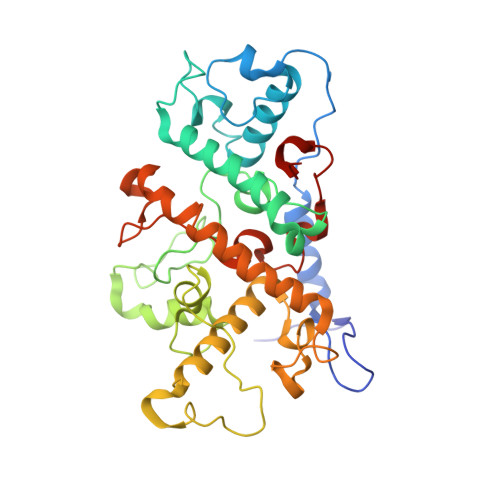
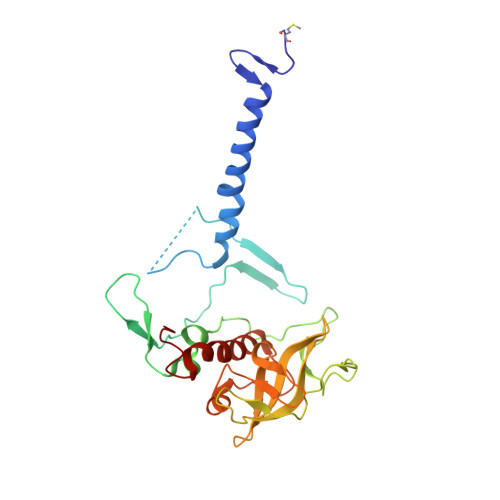
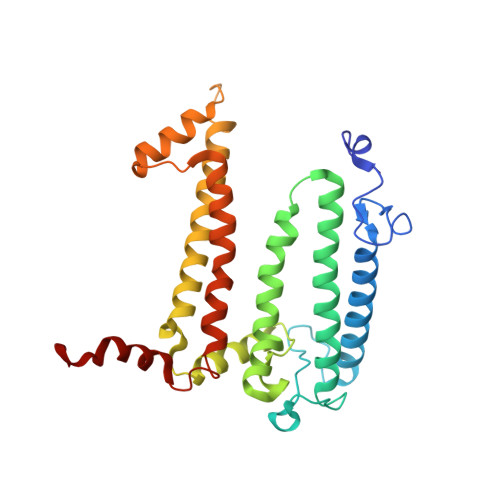
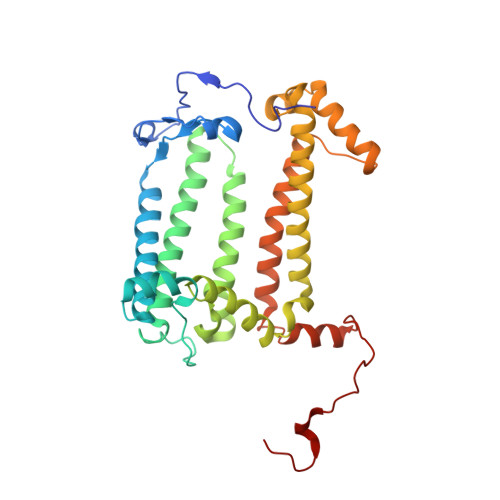
Light-Induced Structural Changes in a Photosynthetic Reaction Center Caught by Laue Diffraction.
Wohri, A.B., Katona, G., Johansson, L.C., Fritz, E., Malmerberg, E., Andersson, M., Vincent, J., Eklund, M., Cammarata, M., Wulff, M., Davidsson, J., Groenhof, G., Neutze, R.(2010) Science 328: 630
- PubMed: 20431017
- DOI: https://doi.org/10.1126/science.1186159
- Primary Citation of Related Structures:
2X5U, 2X5V - PubMed Abstract:
Photosynthetic reaction centers convert the energy content of light into a transmembrane potential difference and so provide the major pathway for energy input into the biosphere. We applied time-resolved Laue diffraction to study light-induced conformational changes in the photosynthetic reaction center complex of Blastochloris viridis. The side chain of TyrL162, which lies adjacent to the special pair of bacteriochlorophyll molecules that are photooxidized in the primary light conversion event of photosynthesis, was observed to move 1.3 angstroms closer to the special pair after photoactivation. Free energy calculations suggest that this movement results from the deprotonation of this conserved tyrosine residue and provides a mechanism for stabilizing the primary charge separation reactions of photosynthesis.
Organizational Affiliation:
Department of Chemical and Biological Engineering, Chalmers University of Technology, Box 462, SE-40530 Göteborg, Sweden.