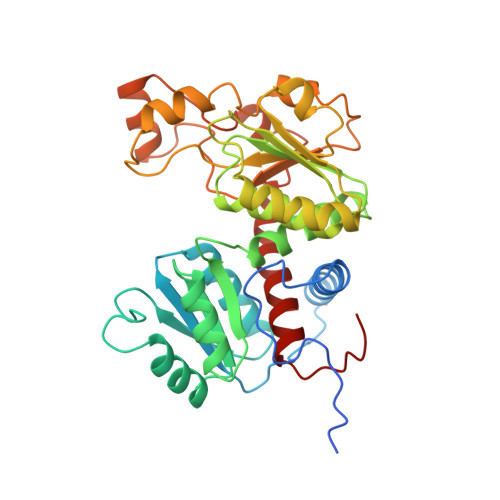
Crystal Structure of the Hexameric Catabolic Ornithine Transcarbamylase from Lactobacillus Hilgardii: Structural Insights Into the Oligomeric Assembly and Metal Binding.
De Las Rivas, B., Fox, G.C., Angulo, I., Ripoll, M.M., Rodriguez, H., Munoz, R., Mancheno, J.M.(2009) J Mol Biol 393: 425
- PubMed: 19666033
- DOI: https://doi.org/10.1016/j.jmb.2009.08.002
- Primary Citation of Related Structures:
2W37 - PubMed Abstract:
Catabolic ornithine transcarbamylase (cOTC; EC 2.1.3.3) catalyzes the formation of ornithine (ORN) and carbamoyl phosphate from citrulline, which constitutes the second step of the degradation of arginine via the arginine deiminase pathway. Here, we report the crystal structure of cOTC from the lactic acid bacteria Lactobacillus hilgardii (Lh-cOTC) refined to 2.1 A resolution. The structure reveals that Lh-cOTC forms a hexameric assembly, which was also confirmed by gel-filtration chromatography and analytical ultracentrifugation. The homohexamer, with 32 point group symmetry, represents a new oligomeric state within the members of the ornithine transcarbamylase family that are typically homotrimeric or homododecameric. The C-terminal end from each subunit constitutes a key structural element for the stabilization of the hexameric assembly in solution. Additionally, the structure reveals, for the first time in the ornithine transcarbamylase family, a metal-binding site located at the 3-fold molecular symmetry axis of each trimer.
Organizational Affiliation:
Departamento de Microbiología, Instituto de Fermentaciones Industriales, CSIC, Madrid, Spain.