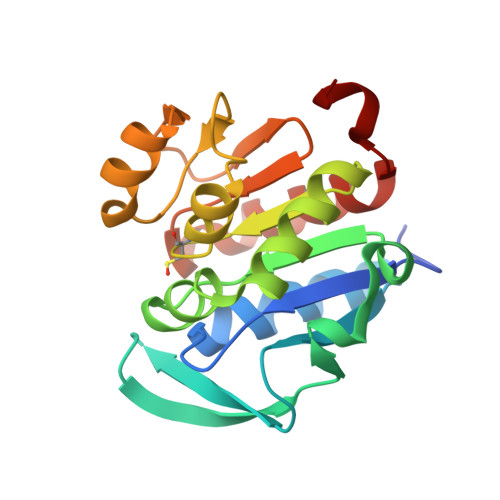
Structure of the stress response protein DR1199 from Deinococcus radiodurans: a member of the DJ-1 superfamily.
Fioravanti, E., Dura, M.A., Lascoux, D., Micossi, E., Franzetti, B., McSweeney, S.(2008) Biochemistry 47: 11581-11589
- PubMed: 18850720
- DOI: https://doi.org/10.1021/bi800882v
- Primary Citation of Related Structures:
2VRN - PubMed Abstract:
The expression level of protein DR1199 is observed to increase considerably in the radio-resistant bacterium Deinococcus radiodurans following irradiation. This protein belongs to the DJ-1 superfamily, which includes proteins with diverse functions, such as the archaeal proteases PhpI and PfpI, the bacterial chaperone Hsp31 and hyperosmotic stress protein YhbO, and the human Parkinson's disease-related protein DJ-1. All members of the superfamily are oligomeric, and the oligomerization interface varies from protein to protein. Although for many of these proteins, their function remains obscure, most of them are involved in cellular protection against environmental stresses. We have determined the structure of DR1199 to a resolution of 2.15 A, and we have tested its function and studied its role in the response to irradiation and more generally to oxidative stress in D. radiodurans. The protein is a dimer displaying an oligomerization interface similar to that observed for the YhbO and PhpI proteins. The cysteine in the catalytic triad (Cys 115) is oxidized in our structure, similar to modifications seen in the corresponding cysteine of the DJ-1 protein. The oxidation occurs spontaneously in DR1199 crystals. In solution, no proteolytic or chaperone activity was detected. On the basis of our results, we suggest that DR1199 might work as a general stress protein involved in the detoxification of the cell from oxygen reactive species, rather than as a peptidase in D. radiodurans.
Organizational Affiliation:
European Synchrotron Radiation Facility, BP 220, 38043 Grenoble Cedex 9, France.