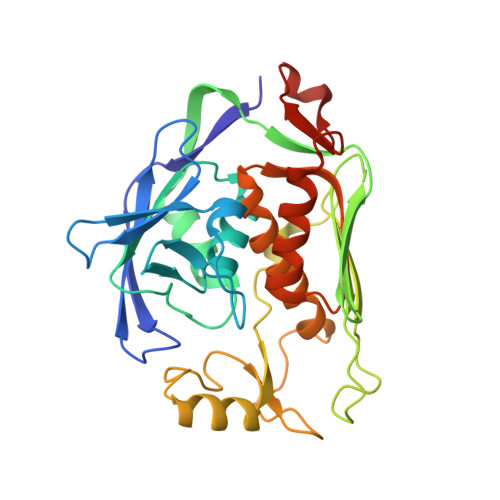
Crystal structure of LpxC from Pseudomonas aeruginosa complexed with the potent BB-78485 inhibitor.
Mochalkin, I., Knafels, J.D., Lightle, S.(2008) Protein Sci 17: 450-457
- PubMed: 18287278
- DOI: https://doi.org/10.1110/ps.073324108
- Primary Citation of Related Structures:
2VES - PubMed Abstract:
The cell wall in Gram-negative bacteria is surrounded by an outer membrane comprised of charged lipopolysaccharide (LPS) molecules that prevent entry of hydrophobic agents into the cell and protect the bacterium from many antibiotics. The hydrophobic anchor of LPS is lipid A, the biosynthesis of which is essential for bacterial growth and viability. UDP-3-O-(R-3-hydroxymyristoyl)-N-acetylglucosamine deacetylase (LpxC) is an essential zinc-dependant enzyme that catalyzes the conversion of UDP-3-O-(R-3-hydroxymyristoyl)-N-acetylglucosamine to UDP-3-O-(R-3-hydroxymyristoyl)glucosamine and acetate in the biosynthesis of lipid A, and for this reason, LpxC is an attractive target for antibacterial drug discovery. Here we disclose a 1.9 A resolution crystal structure of LpxC from Pseudomonas aeruginosa (paLpxC) in a complex with the potent BB-78485 inhibitor. To our knowledge, this is the first crystal structure of LpxC with a small-molecule inhibitor that shows antibacterial activity against a wide range of Gram-negative pathogens. Accordingly, this structure can provide important information for lead optimization and rational design of the effective small-molecule LpxC inhibitors for successful treatment of Gram-negative infections.
Organizational Affiliation:
Pfizer, Inc., Groton, Connecticut 06340, USA. Igor.Mochalkin@pfizer.com