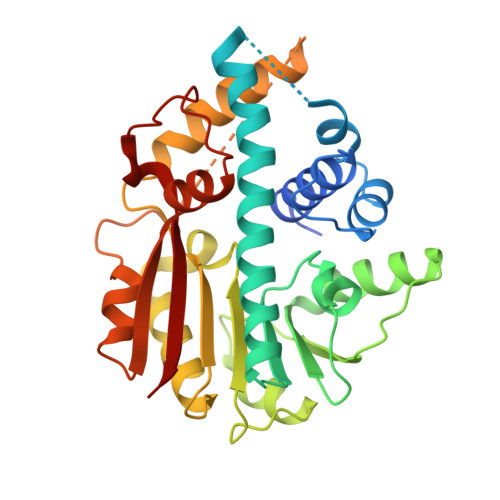
The Crystal Structure of M. Leprae Ml2640C Defines a Large Family of Putative S-Adenosylmethionine- Dependent Methyltransferases in Mycobacteria.
Grana, M., Haouz, A., Buschiazzo, A., Miras, I., Wehenkel, A., Bondet, V., Shepard, W., Schaeffer, F., Cole, S.T., Alzari, P.M.(2007) Protein Sci 16: 1896
- PubMed: 17660248
- DOI: https://doi.org/10.1110/ps.072982707
- Primary Citation of Related Structures:
2CKD, 2UYO, 2UYQ - PubMed Abstract:
Mycobacterium leprae protein ML2640c belongs to a large family of conserved hypothetical proteins predominantly found in mycobacteria, some of them predicted as putative S-adenosylmethionine (AdoMet)-dependent methyltransferases (MTase). As part of a Structural Genomics initiative on conserved hypothetical proteins in pathogenic mycobacteria, we have determined the structure of ML2640c in two distinct crystal forms. As expected, ML2640c has a typical MTase core domain and binds the methyl donor substrate AdoMet in a manner consistent with other known members of this structural family. The putative acceptor substrate-binding site of ML2640c is a large internal cavity, mostly lined by aromatic and aliphatic side-chain residues, suggesting that a lipid-like molecule might be targeted for catalysis. A flap segment (residues 222-256), which isolates the binding site from the bulk solvent and is highly mobile in the crystal structures, could serve as a gateway to allow substrate entry and product release. The multiple sequence alignment of ML2640c-like proteins revealed that the central alpha/beta core and the AdoMet-binding site are very well conserved within the family. However, the amino acid positions defining the binding site for the acceptor substrate display a higher variability, suggestive of distinct acceptor substrate specificities. The ML2640c crystal structures offer the first structural glimpses at this important family of mycobacterial proteins and lend strong support to their functional assignment as AdoMet-dependent methyltransferases.
Organizational Affiliation:
Unité de Biochimie Structurale (CNRS-URA 2185), Institut Pasteur, 75724 Paris, France.