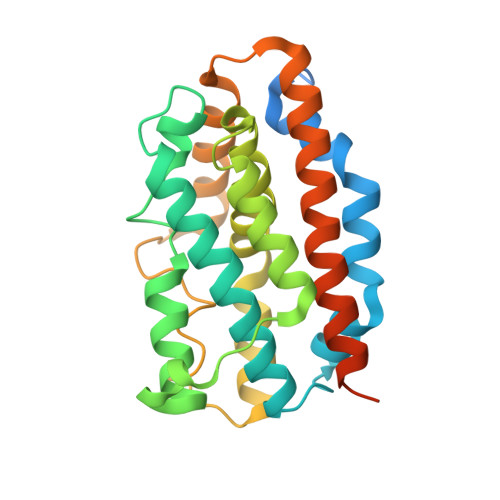
Comparison of Apo- and Heme-bound Crystal Structures of a Truncated Human Heme Oxygenase-2.
Bianchetti, C.M., Yi, L., Ragsdale, S.W., Phillips Jr., G.N.(2007) J Biol Chem 282: 37624-37631
- PubMed: 17965015
- DOI: https://doi.org/10.1074/jbc.M707396200
- Primary Citation of Related Structures:
2Q32, 2QPP, 2RGZ - PubMed Abstract:
Heme oxygenase (HO) catalyzes the first step in the heme degradation pathway. The crystal structures of apo- and heme-bound truncated human HO-2 reveal a primarily alpha-helical architecture similar to that of human HO-1 and other known HOs. Proper orientation of heme in HO-2 is required for the regioselective oxidation of the alpha-mesocarbon. This is accomplished by interactions within the heme binding pocket, which is made up of two helices. The iron coordinating residue, His(45), resides on the proximal helix. The distal helix contains highly conserved glycine residues that allow the helix to flex and interact with the bound heme. Tyr(154), Lys(199), and Arg(203) orient the heme through direct interactions with the heme propionates. The rearrangements of side chains in heme-bound HO-2 compared with apoHO-2 further elucidate HO-2 heme interactions.
Organizational Affiliation:
Graduate Program in Biophysics, Center for Eukaryotic Structural Genomics, University of Wisconsin, Madison, WI 53706, USA.