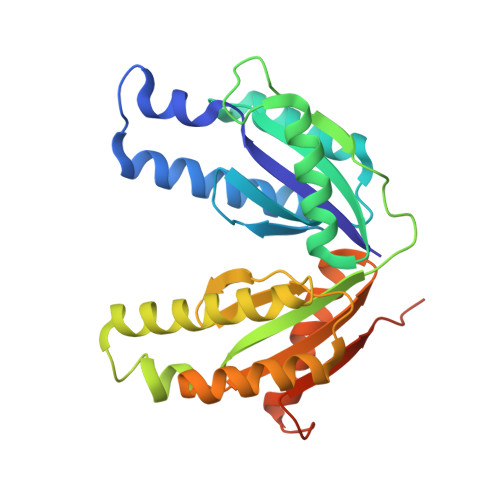
A New Use for a Familiar Fold: The X-ray Crystal Structure of GTP-Bound GTP Cyclohydrolase III from Methanocaldococcus jannaschii Reveals a Two Metal Ion Catalytic Mechanism
Morrison, S.D., Roberts, S.A., Zegeer, A.M., Montfort, W.R., Bandarian, V.(2008) Biochemistry 47: 230-242
- PubMed: 18052207
- DOI: https://doi.org/10.1021/bi701782e
- Primary Citation of Related Structures:
2QV6 - PubMed Abstract:
GTP cyclohydrolase (GCH) III from Methanocaldococcus jannaschii, which catalyzes the conversion of GTP to 2-amino-5-formylamino-6-ribosylamino-4(3H)-pyrimidinone 5'-phosphate (FAPy), has been shown to require Mg2+ for catalytic activity and is activated by monovalent cations such as K+ and ammonium [Graham, D. E., Xu, H., and White, R. H. (2002) Biochemistry 41, 15074-15084]. The reaction is formally identical to that catalyzed by a GCH II ortholog (SCO 6655) from Streptomyces coelicolor; however, SCO 6655, like other GCH II proteins, is a zinc-containing protein. The structure of GCH III complexed with GTP solved at 2 A resolution clearly shows that GCH III adopts a distinct fold that is closely related to the palm domains of phosphodiesterases, such as DNA polymerase I. GCH III is a tetramer of identical subunits; each monomer is composed of an N- and a C-terminal domain that adopt nearly superimposible structures, suggesting that the protein has arisen by gene duplication. Three metal ions were located in the active site, two of which occupy positions that are analogous to those occupied by divalent metal ions in the structures of a number of palm domain containing proteins, such as DNA polymerase I. Two conserved Asp residues that coordinate the metal ions, which are also found in palm domain containing proteins, are observed in GCH III. Site-directed variants (Asp-->Asn) of these residues in GCH III are less active than wild-type. The third metal ion, most likely a potassium ion, is involved in substrate recognition through coordination of O6 of GTP. The arrangement of the metal ions in the active site suggests that GCH III utilizes two metal ion catalysis. The structure of GCH III extends the repertoire of possible reactions with a palm fold to include cyclohydrolase chemistry.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, The University of Arizona, Tucson, Arizona 85721, USA.